User:Ramiro Barrantes/FpgNeiRepair
From Proteopedia
(Difference between revisions)
(→Site-directed mutants) |
(→Site-directed mutants) |
||
| Line 120: | Line 120: | ||
|- | |- | ||
| <scene name='User:Ramiro_Barrantes/Workbench/Ecofpgzincfinger/1' target="ecoNeiMutants">ecoFpg C244(S/H,A)</scene> || E. Coli ([[1k82]]) || No binding nor cleavage || <ref>PMID:8253809</ref> | | <scene name='User:Ramiro_Barrantes/Workbench/Ecofpgzincfinger/1' target="ecoNeiMutants">ecoFpg C244(S/H,A)</scene> || E. Coli ([[1k82]]) || No binding nor cleavage || <ref>PMID:8253809</ref> | ||
| - | | <scene name='User:Ramiro_Barrantes/Workbench/ | + | | <scene name='User:Ramiro_Barrantes/Workbench/Ecofpgzincfingerwithcys247244/1' target="ecoNeiMutants">ecoFpg C244S/C247S)</scene> || E. Coli ([[1k82]]) || No binding nor cleavage. No zinc, no altered secondary structure || <ref>PMID:8253809</ref> |
|} | |} | ||
| <applet load='1K3W' size='500' frame='true' align='left' caption='E. Coli Nei' name="ecoNeiMutants" scene='User:Ramiro_Barrantes/Workbench/Econeiwater/1' /> | | <applet load='1K3W' size='500' frame='true' align='left' caption='E. Coli Nei' name="ecoNeiMutants" scene='User:Ramiro_Barrantes/Workbench/Econeiwater/1' /> | ||
Revision as of 15:21, 9 June 2009
The FpgNei Protein Superfamily
Functional Units
| G. Stereothermophilus Fpg | E. Coli Nei | ||||||||||||
|
|
| Functional Cluster | Variant 1 | Variant 2 | Fpg1 | Fpg2 | Plant | Neil1 | Neil2 | Neil3 | Proteo | Actino1 | Actino2 | MimiVirus |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Support for perfectly conserved Asn168 | Y | Y | N | N | N | N | N | N | N | N | ||
| Stability of catalytic helix | Y | Y | Y | N | N | N | N | N | N | N | ||
| Stability of intercalation loop | Y | Y | Y | N | N | N | N | N | N | Y | ||
| Intercalation loop | Y | Y | Y | N | N | N | N | N | N | Y | ||
| Zinc Finger | Y | Y | N | N | Y | Y | Y | Y | Y | N | ||
| Recognition complex | none | Y | N | N | N | N | N | N | N | N | N | |
| Neil1-specific: Support for Lys60 | Y | Y | Y | N | Y | Y | Y | Y | Y | Y | ||
| PlantFungi-specific: R254 DNA binding | different in plants | Y | Y | N | Y | Y | Y | N | Y | Y | Y |
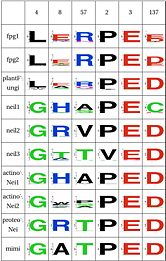
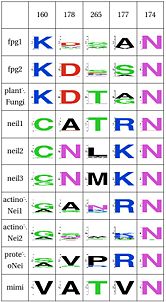
Functional Cluster: Stability of perfectly conserved Asn168Asn74, along with two other amino acids have an effect in the orientation and kinking of the DNA. In 4 of the 9 clades (Fpg1, Fpg2 and Plants and Fungi) Asn174 is supported by Lys160, which in turn hydrogen bonds with Leu249 and Ser250. In the other clades (Actinobacteria 1 and 2, Proteacteria and all vertebrate sequences), Arg171 that comes from a different helix fulfills the same roles as Lys160. One important difference is that the Zinc Finger is shaped differently in the absence of DNA, and there is a hydrogen bond between one of the beta-sheets and the arginine. One hypothesis is that the arginine or the lysine is necessary to support the Asn174, crucial for orientation of the DNA. |
Functional Cluster: Stability of catalytic helixThe triad Leu4, Glu8 and Arg57 interact and provide stability to helixA, which has the catalytic residue Pro2,Glu3 and Glu6. This triad is present in the same four clades as above (Fpg1, Fpg2 and Plants and Fungi). This triad is not present in the remaining clades and it is not clear how the same stability is provided. Leu211 also has a hydrophobic interaction with Leu4. |
Functional Cluster: Stability of intercalation loopImage:IntercalationLoopSupport.jpg This structure provides stability for the amino acids that insert into the space vacated by the damaged base |
Functional Cluster: Stability of key Gly59 and Lys60Gly59 and Lys60 are important in the activity of MutM. We hypothesize that Glu137 is very important to maintain its stability, this amino acid is compensated by Asn172 in Neil1 </ref> |
Functional Cluster: Intercalation LoopImage:IntercalationLoop.jpg The residue in positions 77 and 78 suggest a possible intercalation loop The residue E2 and E6 have been mutated, with the first one inactivating the protein and the second one having no major effect [1]. |
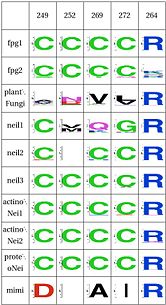
Functional Cluster: Zinc/zincless fingerThe Zinc finger serves to support the Arg264 which binds to the phosphate of the damaged base. In Neil1, there is no Zinc but there is an equivalent structure [2]. Both the plants and mimivirus have a zincless finger, although it is not clear if this one is homologous |
Functional Cluster: Recognition ComplexThis complex is key in recognizing a damaged guanine[3] |
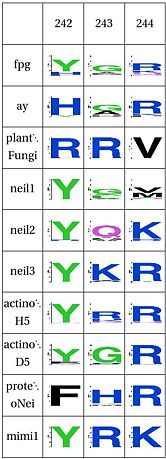
Functional Cluster: DNA binding Tyrosine |
Zinc/zincless finger
Evolution
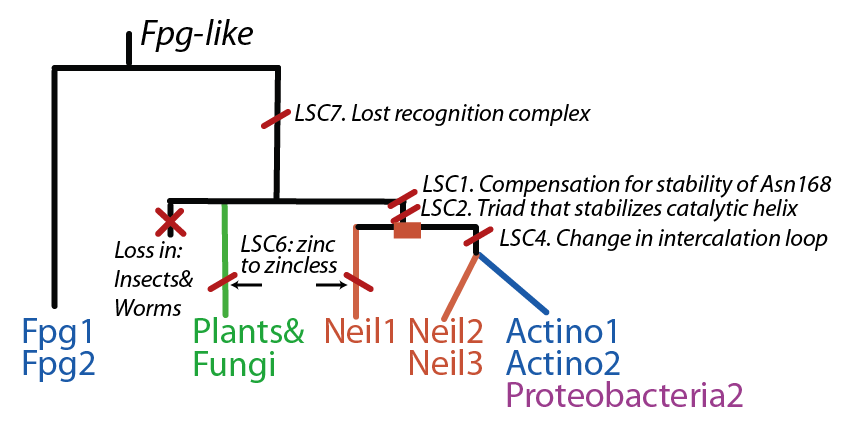
The FpgNei evolution has not been easy to resolve [4], especially in the deeper branches. Assuming that functional clusters evolve more slowly than individual residues, we can use this as phylogenetic characters to 1) draw the most parsimonious evolution of the superfamily as dictated by these functional clusters 2) examine how these clusters have evolved and how this might have influenced the evolution of FpgNei.
Site-directed mutants
|
|
References
- ↑ Burgess S, Jaruga P, Dodson ML, Dizdaroglu M, Lloyd RS. Determination of active site residues in Escherichia coli endonuclease VIII. J Biol Chem. 2002 Jan 25;277(4):2938-44. Epub 2001 Nov 15. PMID:11711552 doi:10.1074/jbc.M110499200
- ↑ Doublie S, Bandaru V, Bond JP, Wallace SS. The crystal structure of human endonuclease VIII-like 1 (NEIL1) reveals a zincless finger motif required for glycosylase activity. Proc Natl Acad Sci U S A. 2004 Jul 13;101(28):10284-9. Epub 2004 Jul 1. PMID:15232006 doi:10.1073/pnas.0402051101
- ↑ Fromme JC, Verdine GL. DNA lesion recognition by the bacterial repair enzyme MutM. J Biol Chem. 2003 Dec 19;278(51):51543-8. Epub 2003 Oct 1. PMID:14525999 doi:10.1074/jbc.M307768200
- ↑ Doublie S, Bandaru V, Bond JP, Wallace SS. The crystal structure of human endonuclease VIII-like 1 (NEIL1) reveals a zincless finger motif required for glycosylase activity. Proc Natl Acad Sci U S A. 2004 Jul 13;101(28):10284-9. Epub 2004 Jul 1. PMID:15232006 doi:10.1073/pnas.0402051101
- ↑ Golan G, Zharkov DO, Feinberg H, Fernandes AS, Zaika EI, Kycia JH, Grollman AP, Shoham G. Structure of the uncomplexed DNA repair enzyme endonuclease VIII indicates significant interdomain flexibility. Nucleic Acids Res. 2005 Sep 6;33(15):5006-16. Print 2005. PMID:16145054 doi:http://dx.doi.org/33/15/5006
- ↑ Sidorkina OM, Laval J. Role of lysine-57 in the catalytic activities of Escherichia coli formamidopyrimidine-DNA glycosylase (Fpg protein). Nucleic Acids Res. 1998 Dec 1;26(23):5351-7. PMID:9826758
- ↑ Golan G, Zharkov DO, Feinberg H, Fernandes AS, Zaika EI, Kycia JH, Grollman AP, Shoham G. Structure of the uncomplexed DNA repair enzyme endonuclease VIII indicates significant interdomain flexibility. Nucleic Acids Res. 2005 Sep 6;33(15):5006-16. Print 2005. PMID:16145054 doi:http://dx.doi.org/33/15/5006
- ↑ Burgess S, Jaruga P, Dodson ML, Dizdaroglu M, Lloyd RS. Determination of active site residues in Escherichia coli endonuclease VIII. J Biol Chem. 2002 Jan 25;277(4):2938-44. Epub 2001 Nov 15. PMID:11711552 doi:10.1074/jbc.M110499200
- ↑ Burgess S, Jaruga P, Dodson ML, Dizdaroglu M, Lloyd RS. Determination of active site residues in Escherichia coli endonuclease VIII. J Biol Chem. 2002 Jan 25;277(4):2938-44. Epub 2001 Nov 15. PMID:11711552 doi:10.1074/jbc.M110499200
- ↑ Burgess S, Jaruga P, Dodson ML, Dizdaroglu M, Lloyd RS. Determination of active site residues in Escherichia coli endonuclease VIII. J Biol Chem. 2002 Jan 25;277(4):2938-44. Epub 2001 Nov 15. PMID:11711552 doi:10.1074/jbc.M110499200
- ↑ Shinmura K, Tao H, Goto M, Igarashi H, Taniguchi T, Maekawa M, Takezaki T, Sugimura H. Inactivating mutations of the human base excision repair gene NEIL1 in gastric cancer. Carcinogenesis. 2004 Dec;25(12):2311-7. Epub 2004 Aug 19. PMID:15319300 doi:10.1093/carcin/bgh267
- ↑ Tchou J, Michaels ML, Miller JH, Grollman AP. Function of the zinc finger in Escherichia coli Fpg protein. J Biol Chem. 1993 Dec 15;268(35):26738-44. PMID:8253809
- ↑ Tchou J, Michaels ML, Miller JH, Grollman AP. Function of the zinc finger in Escherichia coli Fpg protein. J Biol Chem. 1993 Dec 15;268(35):26738-44. PMID:8253809