User:Tilman Schirmer/Sandbox 211
From Proteopedia
(Difference between revisions)
(→Substrate binding) |
(→Substrate binding) |
||
| Line 32: | Line 32: | ||
<br><br><br><br><br><br><br><br><br><br><br><br><br> | <br><br><br><br><br><br><br><br><br><br><br><br><br> | ||
| + | ===Complete active site formed by GGDEF dimer=== | ||
| - | <scene name='User:Tilman_Schirmer/Sandbox_211/Competent_ggdef_dimer/1'>competent GGDEF dimer</scene> | + | |
| + | <scene name='User:Tilman_Schirmer/Sandbox_211/Competent_ggdef_dimer/2'>competent GGDEF dimer 1</scene> | ||
| + | <scene name='User:Tilman_Schirmer/Sandbox_211/Competent_ggdef_dimer/3'>competent GGDEF dimer 2</scene> | ||
| + | <scene name='User:Tilman_Schirmer/Sandbox_211/Competent_ggdef_dimer/1'>competent GGDEF dimer 3</scene> | ||
Revision as of 10:53, 21 June 2009
Overview
|
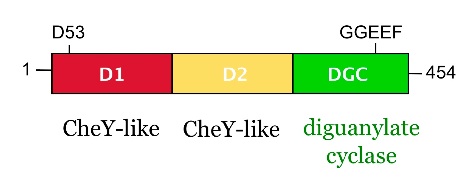
from Caulobacter crescentus is a response regulator with an unorthodox catalytic, diguanylate cyclase, output domain. It is composed of a canonical CheY-like response regulator receiver () domain,
a Rec-like () adaptor domain,
and a C-terminal domain that confers the catalytic acitvity.
The GGDEF domain is named after the highly conserved (in PleD it is GGEEF) that locates to a β-hairpin.
Substrate binding
|
The motif is part of the as identified in the structure of PleD in complex with . The GGDEFY domain binds only one GTP subsrate molecule. For the reaction to proceed, two GTP loaded GGDEF domains have to align antiparallely. MODEL.
Complete active site formed by GGDEF dimer