This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 348
From Proteopedia
| This Sandbox is Reserved from January 10, 2010, through April 10, 2011 for use in BCMB 307-Proteins course taught by Andrea Gorrell at the University of Northern British Columbia, Prince George, BC, Canada. |
To get started:
More help: Help:Editing |
| |||||||||
| 1ppb, resolution 1.92Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Activity: | Thrombin, with EC number 3.4.21.5 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
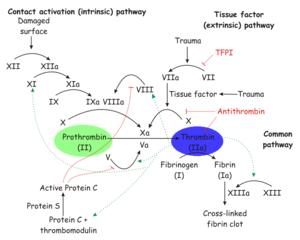
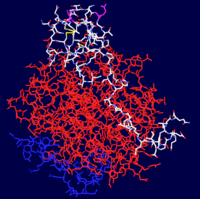
Thrombin is a trypsin-like serine protease which is best known for its role in blood clotting. In humans, the F2 gene codes for prothrombin, which is also known as Coagulation Factor II.[1][2] Clevage of prothrombin to form activated α-thrombin is a key step in the final common pathway of blood clotting, because clevage by thrombin activates several factors in blood clotting, especially fibrin, factor XIII, and protein C.[3]
Contents |
Structure
Thrombin is comprised of two chains, often referred to as the and the . All known functional epitopes are found on the long chain. There is one active site, which in the case of 1ppb is occupied with .[4] Additionally, there are three structural disulfide bonds.
| While thrombin is described as a trypsin-like serine protease, it is more specific than trypsin due to two exosites which bind the substrate at a point separate from the active site. These exosites also allow for more specific inhibition, since protein C and factor Xa have similar active sites but play very different roles in the clotting process.[3] | 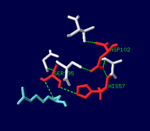 The serine protease catalytic triad in the active site of α-thrombin, bound to D-Phe-Pro-Arg chloromethylketone ligand. |
Regulation
| Prothrombin is proteolytically activated to α-thrombin by factor Xa. α-Thrombin is permanently inactivated by the Serine Protease Inhibitor antithrombin, with heparin as a cofactor, and allosterically regulated by sodium ion concentration. Thrombomodulin inhibits clevage of fibrinogen to fibrin, but also enhances α-thrombin activity with respect to protein C. Since activated protein C proteolytically inactivates earlier steps in the chain, this effectively reverses the role of thrombin from coagulant to anticoagulant.[3]
Hirudin is a potent natural inhibitor of thrombin, produced by leeches such as Hirudo medicinalis. |
3D Structures
α-Thrombin
Prothrombin
See Also
External Resources
- Thrombin at Wikipedia
- Serine protease at Wikipedia
- Fibrin Glue at Wikipedia
- Coagulation (blood clotting) at Wikipedia
- Hemophilia at Wikipedia
References
- ↑ Royle NJ, Irwin DM, Koschinsky ML, MacGillivray RT, Hamerton JL. Human genes encoding prothrombin and ceruloplasmin map to 11p11-q12 and 3q21-24, respectively. Somat Cell Mol Genet. 1987 May;13(3):285-92. PMID:3474786
- ↑ Degen SJ, Davie EW. Nucleotide sequence of the gene for human prothrombin. Biochemistry. 1987 Sep 22;26(19):6165-77. PMID:2825773
- ↑ 3.0 3.1 3.2 Di Cera E. Thrombin interactions. Chest. 2003 Sep;124(3 Suppl):11S-7S. PMID:12970119
- ↑ Bode W, Mayr I, Baumann U, Huber R, Stone SR, Hofsteenge J. The refined 1.9 A crystal structure of human alpha-thrombin: interaction with D-Phe-Pro-Arg chloromethylketone and significance of the Tyr-Pro-Pro-Trp insertion segment. EMBO J. 1989 Nov;8(11):3467-75. PMID:2583108