This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 497
From Proteopedia
| Line 18: | Line 18: | ||
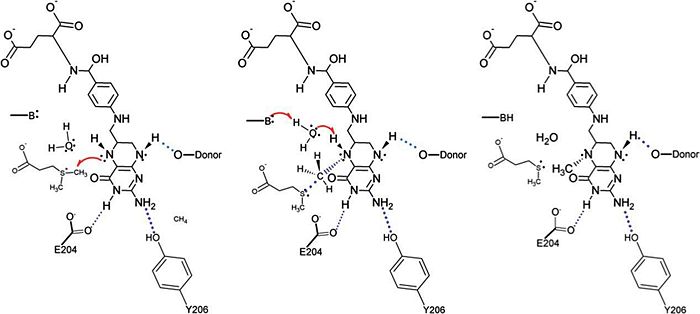
The specific mechanism of DmdA is still being investigated. However, a mechanism was recently proposed <ref> Schuller, D.J., Reisch, C.R., Moran, M.A., Whitman, W.B., Lanzilotta, W.N. (2012) Structures of dimethylsulfoniopropinate-dependent demethylase from the marine organism pelagabacter ubique. Protein Sci. 21: 289-298. </ref> | The specific mechanism of DmdA is still being investigated. However, a mechanism was recently proposed <ref> Schuller, D.J., Reisch, C.R., Moran, M.A., Whitman, W.B., Lanzilotta, W.N. (2012) Structures of dimethylsulfoniopropinate-dependent demethylase from the marine organism pelagabacter ubique. Protein Sci. 21: 289-298. </ref> | ||
| - | [[Image:DmdA_Mechanism.jpg|thumb| | + | [[Image:DmdA_Mechanism.jpg|thumb|700px|left|Proposed mechanism]] |
==Possible Applications== | ==Possible Applications== | ||
Revision as of 16:34, 1 May 2012
| This Sandbox is Reserved from 13/03/2012, through 01/06/2012 for use in the course "Proteins and Molecular Mechanisms" taught by Robert B. Rose at the North Carolina State University, Raleigh, NC USA. This reservation includes Sandbox Reserved 451 through Sandbox Reserved 500. | |||||||
To get started:
More help: Help:Editing For more help, look at this link: http://www.proteopedia.org/wiki/index.php/Help:Getting_Started_in_Proteopedia
Dimethylsulfoniopropionate-Dependent Demethylase A (DmdA))
IntroductionDimethylsulfoniopropionate-Dependendent Demethylase A (DmdA) is an enzyme produced by marine bacterioplankton in order to demethylate Dimethylsulfoniopropionate (DMSP). StructureThe structure of DmdA was recently solved through X-Ray Diffraction [1]. The structure contains 369 amino acids folded into four domains and containing two ligands. Mechanism of ActionThe specific mechanism of DmdA is still being investigated. However, a mechanism was recently proposed [2] Possible ApplicationsReferences
|