This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1gpa
From Proteopedia
(Difference between revisions)
(New page: 200px <!-- The line below this paragraph, containing "STRUCTURE_1gpa", creates the "Structure Box" on the page. You may change the PDB parameter (which sets the PD...) |
m (Protected "1gpa" [edit=sysop:move=sysop]) |
Revision as of 13:41, 3 June 2012
| |||||||||
| 1gpa, resolution 2.90Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||||
| Non-Standard Residues: | |||||||||
| Activity: | Phosphorylase, with EC number 2.4.1.1 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
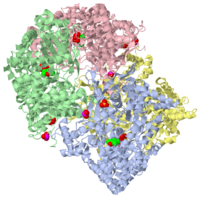
STRUCTURAL MECHANISM FOR GLYCOGEN PHOSPHORYLASE CONTROL BY PHOSPHORYLATION AND AMP
Template:ABSTRACT PUBMED 1900534
About this Structure
1gpa is a 4 chain structure of Glycogen Phosphorylase with sequence from Oryctolagus cuniculus. The December 2001 RCSB PDB Molecule of the Month feature on Glycogen Phosphorylase by David S. Goodsell is 10.2210/rcsb_pdb/mom_2001_12. Full crystallographic information is available from OCA.
See Also
Reference
- Barford D, Hu SH, Johnson LN. Structural mechanism for glycogen phosphorylase control by phosphorylation and AMP. J Mol Biol. 1991 Mar 5;218(1):233-60. PMID:1900534
- Oikonomakos NG, Tsitsanou KE, Zographos SE, Skamnaki VT, Goldmann S, Bischoff H. Allosteric inhibition of glycogen phosphorylase a by the potential antidiabetic drug 3-isopropyl 4-(2-chlorophenyl)-1,4-dihydro-1-ethyl-2-methyl-pyridine-3,5,6-tricarbo xylate. Protein Sci. 1999 Oct;8(10):1930-45. PMID:10548038