This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox reserved 916
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 3: | Line 3: | ||
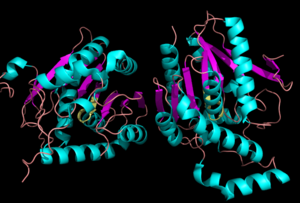
[[Image:Complete_crystal_structure.png|left|300px|thumb|Crystal Structure of MGL]] | [[Image:Complete_crystal_structure.png|left|300px|thumb|Crystal Structure of MGL]] | ||
==Background== | ==Background== | ||
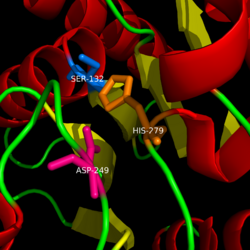
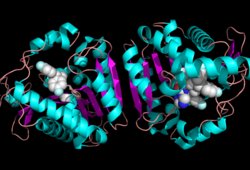
| - | Monoglyceride lipase is part of the α/β hydrolase family, having a Ser-His-Asp catalytic triad (Celemnte et al. 2012). This enzyme is present in most cells, providing the rate limiting step for MG (Taschler et al 2011). MGL terminates the signaling of a primary endocannabinoid, 2-AG (Savinainen et al 2010). MGL is | + | Monoglyceride lipase is part of the α/β hydrolase family, having a Ser-His-Asp catalytic triad (Celemnte et al. 2012). This enzyme is present in most cells, providing the rate limiting step for MG (Taschler et al 2011). MGL terminates the signaling of a primary endocannabinoid, 2-AG (Savinainen et al 2010). MGL is the main enzyme respondsible for hydrolyzing 2-arachidonoylglycerol into arachidonic acid and glycerol ''in vivo'' (Bertrand et al. 2010). One of the key features of MGL is the hydrophobic tunnel, which has been suggested to provide a model for drug research. |
===Metabolic Role=== | ===Metabolic Role=== | ||
| Line 35: | Line 35: | ||
== References == | == References == | ||
| - | Savinainen, Juha R., Megumi Yoshino, Anna Minkkilä, Tapio Nevalainen, and Jarmo T. Laitinen. "Characterization of Binding Properties of Monoglyceride Lipase Inhibitors by a Versatile Fluorescence-based Technique." Analytical Biochemistry 399.1 (2010): 132-34 | + | -Clemente, J. C., E. Nulton, M. Nelen, M. J. Todd, D. Maguire, C. Schalk-Hihi, L. C. Kuo, S.-P. Zhang, C. M. Flores, and J. K. Kranz. "Screening and Characterization of Human Monoglyceride Lipase Active Site Inhibitors Using Orthogonal Binding and Functional Assays." Journal of Biomolecular Screening 17.5 (2012): 629-40. |
| + | |||
| + | -Savinainen, Juha R., Megumi Yoshino, Anna Minkkilä, Tapio Nevalainen, and Jarmo T. Laitinen. "Characterization of Binding Properties of Monoglyceride Lipase Inhibitors by a Versatile Fluorescence-based Technique." Analytical Biochemistry 399.1 (2010): 132-34 | ||
| + | |||
| + | -Taschler, U., F. P. W. Radner, C. Heier, R. Schreiber, M. Schweiger, G. Schoiswohl, K. Preiss-Landl, D. Jaeger, B. Reiter, H. C. Koefeler, J. Wojciechowski, C. Theussl, J. M. Penninger, A. Lass, G. Haemmerle, R. Zechner, and R. Zimmermann. "Monoglyceride Lipase Deficiency in Mice Impairs Lipolysis and Attenuates Diet-induced Insulin Resistance." Journal of Biological Chemistry 286.20 (2011): 17467-7477 | ||
<references/> | <references/> | ||
Current revision
Monoglyceride Lipase (MGL)
| |||||||||||