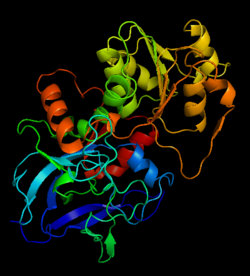
Alcohol dehydrogenase
From Proteopedia
| Line 22: | Line 22: | ||
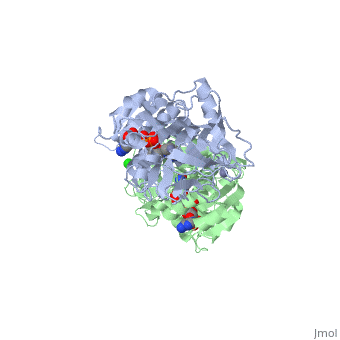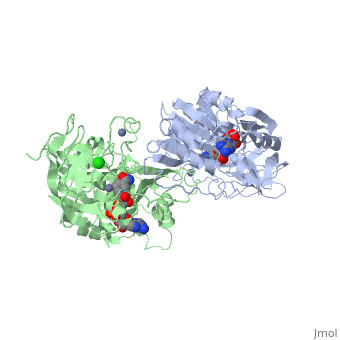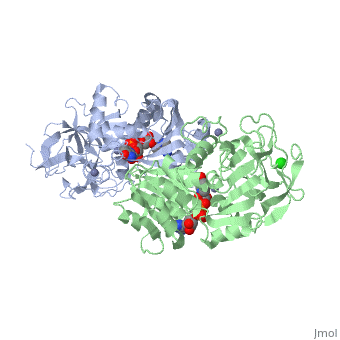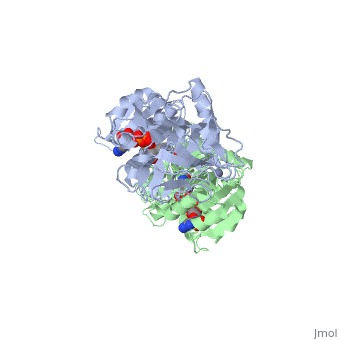
The <scene name='3fsr/Al1/2'>comparison</scene> of overall Cα backbone of all these chimeras (rmsd 0.45-0.65 Å) with those of the parent enzymes, did not reveal significant structural changes. So, the differences in the thermal stability of the chimeras and the parent enzymes could be caused by relatively small specific changes located at the important points of the NADP<sup>+</sup>-dependent alcohol dehydrogenases. For example see Cα superposition for the <font color='red'><b>X23<sub>(TET)</sub> chimera (red)</b></font> ([[3fpc]]) and its parent ADHs (<font color='blue'><b>TbADH, colored blue</b></font> ([[1ped]]), and <font color='lime'><b>EhADH1, colored lime</b></font> ([[1y9a]]). The [http://en.wikipedia.org/wiki/Root_mean_square_deviation RMSDs] of the TbADH−EhADH1, TbADH−Χ23<sub>(TET)</sub>, and EhADH1−Χ23<sub>(TET)</sub> were 0.68, 0.56, and 0.48 Å, respectively. | The <scene name='3fsr/Al1/2'>comparison</scene> of overall Cα backbone of all these chimeras (rmsd 0.45-0.65 Å) with those of the parent enzymes, did not reveal significant structural changes. So, the differences in the thermal stability of the chimeras and the parent enzymes could be caused by relatively small specific changes located at the important points of the NADP<sup>+</sup>-dependent alcohol dehydrogenases. For example see Cα superposition for the <font color='red'><b>X23<sub>(TET)</sub> chimera (red)</b></font> ([[3fpc]]) and its parent ADHs (<font color='blue'><b>TbADH, colored blue</b></font> ([[1ped]]), and <font color='lime'><b>EhADH1, colored lime</b></font> ([[1y9a]]). The [http://en.wikipedia.org/wiki/Root_mean_square_deviation RMSDs] of the TbADH−EhADH1, TbADH−Χ23<sub>(TET)</sub>, and EhADH1−Χ23<sub>(TET)</sub> were 0.68, 0.56, and 0.48 Å, respectively. | ||
<br/> | <br/> | ||
| - | {{TOC limit|limit=2}} | ||
{{Clear}} | {{Clear}} | ||
| Line 82: | Line 81: | ||
{{Clear}} | {{Clear}} | ||
</StructureSection> | </StructureSection> | ||
| - | + | ||
==Additional Resources== | ==Additional Resources== | ||
For additional information, see: [[Carbohydrate Metabolism]]<br /> | For additional information, see: [[Carbohydrate Metabolism]]<br /> | ||
| Line 128: | Line 127: | ||
[[1e3i]] - mADH II + NADH + inhibitor | [[1e3i]] - mADH II + NADH + inhibitor | ||
| - | + | ===ADH III=== | |
[[1m6h]], [[1m6w]], [[1teh]] - hADH III χ chain<br /> | [[1m6h]], [[1m6w]], [[1teh]] - hADH III χ chain<br /> | ||
| Line 152: | Line 151: | ||
[[1d1s]], [[1agn]] – hADH IV σ chain + NAD<br /> | [[1d1s]], [[1agn]] – hADH IV σ chain + NAD<br /> | ||
[[1d1t]] - hADH IV σ chain (mutant) + NAD | [[1d1t]] - hADH IV σ chain (mutant) + NAD | ||
| - | |||
''ADH IV ternary complex'' | ''ADH IV ternary complex'' | ||
| Line 283: | Line 281: | ||
[[1e6w]] - rSHCDH + estradiol + NAD<BR /> | [[1e6w]] - rSHCDH + estradiol + NAD<BR /> | ||
| - | + | ===Unspecified HADH=== | |
[[1uay]] - HADH II – ''Thermus thermophilus''<BR /> | [[1uay]] - HADH II – ''Thermus thermophilus''<BR /> | ||
| Line 290: | Line 288: | ||
[[2x58]] - rHADH + CoA <BR /> | [[2x58]] - rHADH + CoA <BR /> | ||
[[2et6]] – HADH (mutant) – ''Candida tropicalis'' | [[2et6]] – HADH (mutant) – ''Candida tropicalis'' | ||
| - | |||
| - | |||
| - | |||
==References== | ==References== | ||
Revision as of 08:55, 14 August 2014
| |||||||||||
Contents |
Additional Resources
For additional information, see: Carbohydrate Metabolism
3D Structures of Alcohol dehydrogenase
Updated on 14-August-2014
ADH I
3jv7 – RrADH I – Rhodococcus rubber
2vna - hADH I catalytic domain - human
2hcy – yADH I – yeast
4eex – LlADH I – Lactococcus lactis
4eez – LlADH I (mutant)
ADH I binary complex
1u3t – hADH I α chain + inhibitor
1hsz, 1hdz, 3hud - hADH I β chain + NAD
1u3w - hADH I γ chain + inhibitor
1ht0 - hADH I γ chain (mutant) + NAD
ADH I ternary complex
2xaa – RrADH I + NAD + alcohol
3fx4 – pADH I + NADP + inhibitor – pig
2w98, 2w4q – hADH I catalytic domain + NADP + inhibitor
1hso - hADH I α chain + NAD + pyrazole derivative
1hdx - hADH I β chain + NAD + alcohol
1u3u, 1u3v - hADH I β chain + inhibitor
1deh, 1hdy - hADH I β chain + NAD + pyrazole derivative
1htb - hADH I β3 chain + NAD + pyrazole derivative
ADH II
3owo – ZmADH II iron-dependent – Zymomonas mobilis
ADH II binary complex
3ox4 - ZmADH II iron-dependent + NAD
3cos - hADH II + NAD + Zn
1e3e – mADH II + NADH – mouse
1e3l - mADH II (mutant) + NADH
1e3i - mADH II + NADH + inhibitor
ADH III
1m6h, 1m6w, 1teh - hADH III χ chain
2fze - hADH III χ chain + ADP-ribose
2fzw, 1mp0 - hADH III χ chain + NAD
1mc5 – hADH III χ chain + glutathione + NADH
1ma0 - hADH III χ chain + dodecanoic acid + NAD
3qj5 - hADH III χ chain + inhibitor + NAD
4dl9, 4dlb – tADH III + NAD – tomato
4dla – tADH III
ADH IV
1ye3, 8adh, 5adh - hoADH IV e chain – horse
1qlj - hoADH IV e chain (mutant)
3iv7 – ADH IV – Corynebacterium glutamicum
ADH IV binary complex
2jhf, 2jhg, 1het, 1heu, 1hf3, 1ee2, 2oxi, 2ohx, 6adh - hoADH IV e chain + NAD
1adb, 1adc, 1adf, 1adg, 7adh - hoADH IV e chain + NAD derivative
1mgo, 1ju9, 1qlh, 1a72 - hoADH IV e chain (mutant) + NAD
1d1s, 1agn – hADH IV σ chain + NAD
1d1t - hADH IV σ chain (mutant) + NAD
ADH IV ternary complex
3oq6, 1qv6, 1qv7, 1a71, 1axe, 1axg, 4nfh, 4nfs, 4ng5 – hoADH IV e chain (mutant) + NAD + alcohol
4dwv, 4dxh - hoADH IV e chain + NAD + alcohol
1p1r, 1ldy, 1lde - hoADH IV e chain + NADH + formamide derivative
1n92 - hoADH IV e chain + NAD + pyrazole derivative
1bto, 3bto - hoADH IV e chain + NADH + butylthiolane derivative
1n8k - hoADH IV e chain (mutant) + NAD + pyrazole
1mg0, 1hld - hoADH IV e chain + NAD + alcohol
ADH
1a4u – SlADH – Scaptodrosophila lebanonensis
3my7 – ADH ACDH domain – Vibrio parahaemolyticus
3meq – ADH – Brucella suis
3l4p – ADH – Desulfovibrio gigas
1jvb - SsADH – Sulfolobus solfataricus
3i4c, 1nto, 1nvg – SsADH (mutant)
3goh – ADH – Shewanella oneidensis
3gaz – ADH residues 2-334 – Novosphingobium aromaticivorans
2eih – ADH – Thermus thermophilus
1rjw – GsADH – Geobacillus stearothermophilus
1vj0, 1vhd – TmADH -Thermotoga maritima
2eer – ADH – Sulfolobus tokodaii
3uog – ADH – Sinorhizobium meliloti
ADH binary complex
3l77, 3tn7 – ADH short-chain + NADP – Thermococcus sibiricus
1h2b – ADH + NAD – Aeropyrum pernix
1f8f – Benzyl-ADH + NAD – Acinetobacter calcoaceticus
1o2d - TmADH + NADP
3ip1 – TmADH + Cd
1b16, 1b14, 1b15 - SlADH + NAD derivative
1cdo – ADH + NAD - cod
1rhc – ADH F420-dependent +F420-acetone – Methanoculleus thermophilus
3s2e – ReADH + NAD + Zn
1agn – hADH (sigma) +NAD
3pii – GsADH + butyramide
3rj5, 3rj9 – SlADH (mutant) + NAD
3s1l – ReADH + Zn – Ralstonia eutropha
3jzd – ReADH + NAD
ADH ternary complex
1mg5 – ADH + NADH + acetate – Drosophila melanogaster
1r37 – SsADH + NAD + alcohol
1sby – SlADH + NAD + alcohol
1b2l - SlADH + NAD + cyclohexanone
1llu - ADH + NAD + alcohol – Pseudomonas aeruginosa
3cv7 – pADH + NAD + NAP
3rf7 – SoADH + NAD + Fe + Ni
3s2e – ReADH + NAD + Zn
3s2f, 3s2g – ReADH + NAD + Zn + furfural
4gkv – ADH + NAD + Zn + peptide – Escherichia coli
4jji, 4gl4, 3uko - AtADH III + NAD + Zn – Arabidopsis thaliana
4l0q - AtADH III (mutant) + NAD + Zn
NADP-dependent ADH
1ped - CbADH – Clostridium beijerinckii
2b83, 1jqb – CbADH (mutant)
2nvb - TbADH (mutant) – Thermoanaerobacter brockii
3ftn, 3fpc, 3fpl, 3fsr – ADH chimera
1y9a - EhADH – Entamoeba histolytica
2oui – EhADH (mutant)
1p0c – RpADH8 – Rana perezi
4hfj – toADH – tobacco
4gac - mADH
NADP-dependent ADH binary complex
1kev – CbADH + NADPH
1bxz – CbADH catalytic domain + alcohol
1ykf – TbADH + NADP
3h4g – pADH + NADP
1p0f – RpADH + NADP
4hfm - toADH + NADP
4hfn - toADH + NADP + coniferaldehyde
4jbg - PaADH + Zn – Pyrobaculum aerophilum
4jbh - PaADH + Zn + Co
4jbi - PaADH + NADP + Zn
R-specific ADH
1nxq - LbRADH – Lactobacillus brevis
1zk2, 1zk3 - LbRADH (mutant)
1zjy, 1zjz, 1zk0, 1zk1 – LbRADH (mutant) + NADH + alcohol
1zk4 - LbRADH (mutant) + NADH + acetophenone
Specific alcohol ADH
2cf5, 2cf6 – Cinnamyl-AtADH
1piw, 1q1n, 1ps0 – Cinnamyl-yADH
3two - Cinnamyl-ADH + NADP – Helicobacter pylori
1m2w – Mannitol-ADH – Pseudomonas fluorescens
1w6s – Methanol-ADH – Methylobacterium extorquens
1yqx – Sinapyl-aADH II – aspen
1yqd – Sinapyl-aADH II + NADP
1bdb – Biphenyl dihydrodiol-ADH + NAD - Pseudomonas
Quinohemoprotein ADH
1kv9, 1yiq – PpQADH II + PQQ + heme – Pseudomonas putida
1kb0 - QADH I + PQQ + heme – Comamonas testosteroni
Hydroxyacyl-CoA dehydrogenase
Short chain HADH
1so8 – hSHCDH II – human
3rqs - hSHCDH
1f14 - hSHCDH (mutant)
Short chain HADH binary complex
1f12 - hSHCDH (mutant) + hydroxybutyryl-CoA
1f17, 1lsj, 1lso - hSHCDH (mutant) + NAD
1zbq - hSHCDH IV + NAD
1e3s - rSHCDH + NAD – rat
Short chain HADH ternary complex
1u7t - hSHCDH II + inhibitor + NAD
1f0y - hSHCDH + acetoacetyl-CoA + NAD
1il0, 1m75, 1m76 - hSHCDH (mutant) + acetoacetyl-CoA + NAD
1e3w - rSHCDH + 3-keto-butyrate + NAD
1e6w - rSHCDH + estradiol + NAD
Unspecified HADH
1uay - HADH II – Thermus thermophilus
1zej, 3ctv - HADH – Archaeoglobus fulgidus
1zcj - rHADH
2x58 - rHADH + CoA
2et6 – HADH (mutant) – Candida tropicalis
References
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol dehydrogenase from Human (Homo sapiens), different isozymes. SCOP. 2009. 1 March 2010 < http://scop.berkeley.edu/data/scop.b.d.c.b.b.c.html>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Protein: Alcohol Dehydrogenase. The College of Saint Benedict and Saint John's University. 1 March 2010 < http://www.users.csbsju.edu/~hjakubow/classes/rasmolchime/99ch331proj/alcoholdehydro/index.htm>
- ↑ Voet, et. al. Fundamentals of Biochemistry: 3rd Edition. Hoboken: Wiley & Sons, Inc, 2008.
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Bille V, Remacle J. Simple-kinetic descriptions of alcohol dehydrogenase after immobilization on tresyl-chloride-activated agarose. Eur J Biochem. 1986 Oct 15;160(2):343-8. PMID:3769934
- ↑ Dickinson FM, Monger GP. A study of the kinetics and mechanism of yeast alcohol dehydrogenase with a variety of substrates. Biochem J. 1973 Feb;131(2):261-70. PMID:4352908
- ↑ Blomstrand R, Ostling-Wintzell H, Lof A, McMartin K, Tolf BR, Hedstrom KG. Pyrazoles as inhibitors of alcohol oxidation and as important tools in alcohol research: an approach to therapy against methanol poisoning. Proc Natl Acad Sci U S A. 1979 Jul;76(7):3499-503. PMID:115004
- ↑ Alcohol Dehydrogenase. Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Alcohol Dehydrogenase.Worthington Biochemical Corporation . 31 March 2010 < http://http://www.worthington-biochem.com/ADH/default.html>
- ↑ Goihberg E, Dym O, Tel-Or S, Levin I, Peretz M, Burstein Y. A single proline substitution is critical for the thermostabilization of Clostridium beijerinckii alcohol dehydrogenase. Proteins. 2007 Jan 1;66(1):196-204. PMID:17063493 doi:10.1002/prot.21170
- ↑ Goihberg E, Dym O, Tel-Or S, Shimon L, Frolow F, Peretz M, Burstein Y. Thermal stabilization of the protozoan Entamoeba histolytica alcohol dehydrogenase by a single proline substitution. Proteins. 2008 Feb 7;. PMID:18260103 doi:10.1002/prot.21946
- ↑ Goihberg E, Peretz M, Tel-Or S, Dym O, Shimon L, Frolow F, Burstein Y. Biochemical and Structural Properties of Chimeras Constructed by Exchange of Cofactor-Binding Domains in Alcohol Dehydrogenases from Thermophilic and Mesophilic Microorganisms. Biochemistry. 2010 Feb 9. PMID:20102159 doi:10.1021/bi901730x
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, David Birrer