This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
3int
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
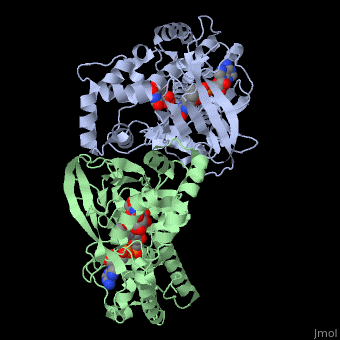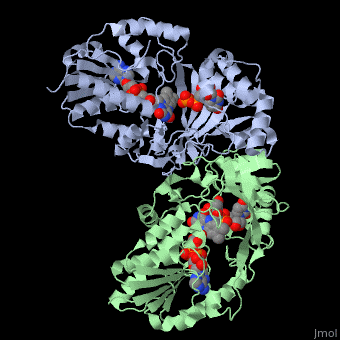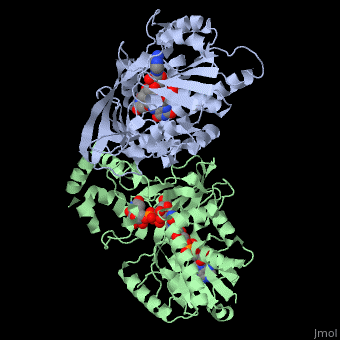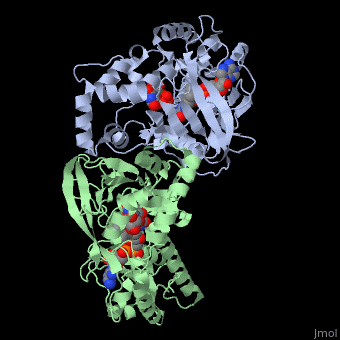
| - | [[ | + | ==Structure of UDP-galactopyranose mutase bound to UDP-galactose (reduced)== |
| + | <StructureSection load='3int' size='340' side='right' caption='[[3int]], [[Resolution|resolution]] 2.51Å' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[3int]] is a 2 chain structure with sequence from [http://en.wikipedia.org/wiki/Klebsiella_pneumoniae Klebsiella pneumoniae]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=3INT OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=3INT FirstGlance]. <br> | ||
| + | </td></tr><tr><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=FDA:DIHYDROFLAVINE-ADENINE+DINUCLEOTIDE'>FDA</scene>, <scene name='pdbligand=GDU:GALACTOSE-URIDINE-5-DIPHOSPHATE'>GDU</scene>, <scene name='pdbligand=UDP:URIDINE-5-DIPHOSPHATE'>UDP</scene><br> | ||
| + | <tr><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat">[[1i8t|1i8t]], [[2bi7|2bi7]], [[2bi8|2bi8]], [[1wam|1wam]], [[1v0j|1v0j]], [[3gf4|3gf4]], [[3inr|3inr]]</td></tr> | ||
| + | <tr><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">glf, rfbD ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=573 Klebsiella pneumoniae])</td></tr> | ||
| + | <tr><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/UDP-galactopyranose_mutase UDP-galactopyranose mutase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.4.99.9 5.4.99.9] </span></td></tr> | ||
| + | <tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=3int FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3int OCA], [http://www.rcsb.org/pdb/explore.do?structureId=3int RCSB], [http://www.ebi.ac.uk/pdbsum/3int PDBsum]</span></td></tr> | ||
| + | <table> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/in/3int_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | The flavoenzyme uridine 5'-diphosphate galactopyranose mutase (UGM or Glf) catalyzes the interconversion of UDP-galactopyranose and UDP-galactofuranose. The latter is a key building block for cell wall construction in numerous pathogens, including Mycobacterium tuberculosis. Mechanistic studies of UGM suggested a novel role for the flavin, and we previously provided evidence that the catalytic mechanism proceeds through a covalent flavin-galactose iminium. Here, we describe 2.3 and 2.5 A resolution X-ray crystal structures of the substrate-bound enzyme in oxidized and reduced forms, respectively. In the latter, C1 of the substrate is 3.6 A from the nucleophilic flavin N5 position. This orientation is consistent with covalent catalysis by flavin. | ||
| - | + | X-ray Crystallography Reveals a Reduced Substrate Complex of UDP-Galactopyranose Mutase Poised for Covalent Catalysis by Flavin .,Gruber TD, Westler WM, Kiessling LL, Forest KT Biochemistry. 2009 Sep 10. PMID:19719175<ref>PMID:19719175</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
==See Also== | ==See Also== | ||
*[[UDP-galactopyranose mutase|UDP-galactopyranose mutase]] | *[[UDP-galactopyranose mutase|UDP-galactopyranose mutase]] | ||
| - | + | == References == | |
| - | == | + | <references/> |
| - | < | + | __TOC__ |
| + | </StructureSection> | ||
[[Category: Klebsiella pneumoniae]] | [[Category: Klebsiella pneumoniae]] | ||
[[Category: UDP-galactopyranose mutase]] | [[Category: UDP-galactopyranose mutase]] | ||
Revision as of 12:34, 29 September 2014
Structure of UDP-galactopyranose mutase bound to UDP-galactose (reduced)
| |||||||||||