This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Clp Protease
From Proteopedia
| Line 1: | Line 1: | ||
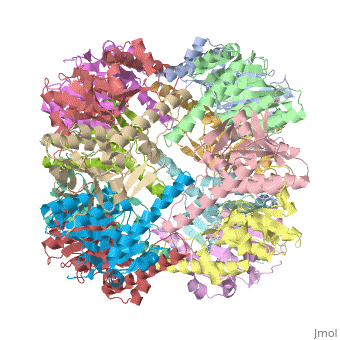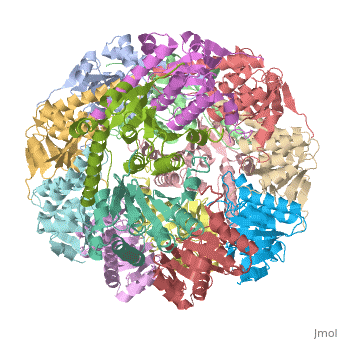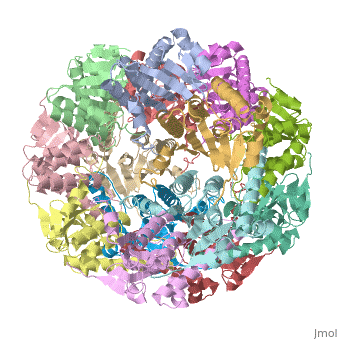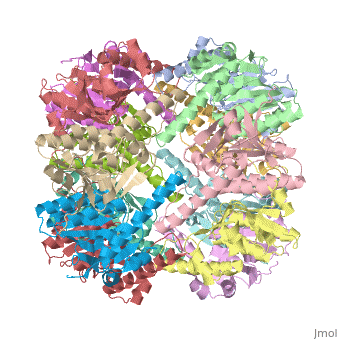
| - | {{STRUCTURE_1tyf| PDB=1tyf | SIZE= | + | {{STRUCTURE_1tyf| PDB=1tyf | SIZE=350| SCENE= |right|CAPTION=E. coli Clp protease, [[1tyf]] }} |
Revision as of 11:23, 8 December 2015
| |||||||||
| E. coli Clp protease, 1tyf | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Activity: | Endopeptidase Clp, with EC number 3.4.21.92 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Clp protease (CLP) is a serine peptidase which hydrolyzes proteins in the presence of ATP and Mg+2 ion. CLP is found in mitochondria. CLP participates in degradation of misfolded proteins. CLP is a heterodimer containing an ATP-binding regulatory subunit A and catalytic subunit P. For more details see
3D structures of Clp protease
Updated on 08-December-2015