This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox flippases
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
==Flippases== | ==Flippases== | ||
<StructureSection load='5c76' size='340' side='right' caption='Caption for this structure' scene=''> | <StructureSection load='5c76' size='340' side='right' caption='Caption for this structure' scene=''> | ||
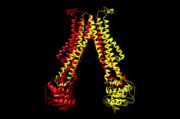
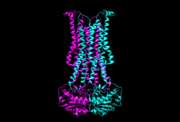
| + | '''Flippases''' are the homodimeric transmembrane proteins located in the membrane. It helps phospholipid to transport from inner face to outer face of the cell membrane. It is composed with alpha-helix and beta sheets. The main three structural features of each monomer are an external helix, a belt of positively charged amino acids, and a binding site for an ATP molecule. | ||
This is a default text for your page '''Sandbox flippases'''. Click above on '''edit this page''' to modify. Be careful with the < and > signs. | This is a default text for your page '''Sandbox flippases'''. Click above on '''edit this page''' to modify. Be careful with the < and > signs. | ||
You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | ||
== Function == | == Function == | ||
| - | + | Flippases flip the lipid from cytoplasmic side to periplasmic side. | |
== Disease == | == Disease == | ||
Revision as of 03:41, 16 December 2015
This page is setup for Leah to build her senior project for OU CHEM 4923
Flippases
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644