This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1iat
From Proteopedia
| Line 4: | Line 4: | ||
|PDB= 1iat |SIZE=350|CAPTION= <scene name='initialview01'>1iat</scene>, resolution 1.62Å | |PDB= 1iat |SIZE=350|CAPTION= <scene name='initialview01'>1iat</scene>, resolution 1.62Å | ||
|SITE= | |SITE= | ||
| - | |LIGAND= <scene name='pdbligand= | + | |LIGAND= <scene name='pdbligand=BME:BETA-MERCAPTOETHANOL'>BME</scene>, <scene name='pdbligand=SO4:SULFATE+ION'>SO4</scene> |
| - | |ACTIVITY= [http://en.wikipedia.org/wiki/Glucose-6-phosphate_isomerase Glucose-6-phosphate isomerase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.3.1.9 5.3.1.9] | + | |ACTIVITY= <span class='plainlinks'>[http://en.wikipedia.org/wiki/Glucose-6-phosphate_isomerase Glucose-6-phosphate isomerase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.3.1.9 5.3.1.9] </span> |
|GENE= GPI ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]) | |GENE= GPI ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]) | ||
| + | |DOMAIN= | ||
| + | |RELATEDENTRY=[[1dqr|1DQR]] | ||
| + | |RESOURCES=<span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1iat FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1iat OCA], [http://www.ebi.ac.uk/pdbsum/1iat PDBsum], [http://www.rcsb.org/pdb/explore.do?structureId=1iat RCSB]</span> | ||
}} | }} | ||
| Line 16: | Line 19: | ||
==Disease== | ==Disease== | ||
| - | Known | + | Known disease associated with this structure: Hemolytic anemia due to glucosephosphate isomerase deficiency OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=172400 172400]], Hydrops fetalis, one form OMIM:[[http://www.ncbi.nlm.nih.gov/entrez/dispomim.cgi?id=172400 172400]] |
==About this Structure== | ==About this Structure== | ||
| Line 32: | Line 35: | ||
[[Category: Pearce, J.]] | [[Category: Pearce, J.]] | ||
[[Category: Read, J A.]] | [[Category: Read, J A.]] | ||
| - | [[Category: BME]] | ||
| - | [[Category: SO4]] | ||
[[Category: glycolysis enzyme/neurotrophic growth factor/cytokine]] | [[Category: glycolysis enzyme/neurotrophic growth factor/cytokine]] | ||
[[Category: isomerase]] | [[Category: isomerase]] | ||
[[Category: two alpha/beta domain]] | [[Category: two alpha/beta domain]] | ||
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Sun Mar 30 21:17:07 2008'' |
Revision as of 18:17, 30 March 2008
| |||||||
| , resolution 1.62Å | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||
| Gene: | GPI (Homo sapiens) | ||||||
| Activity: | Glucose-6-phosphate isomerase, with EC number 5.3.1.9 | ||||||
| Related: | 1DQR
| ||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
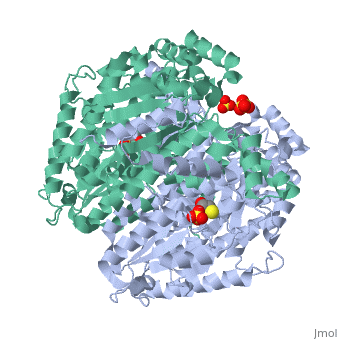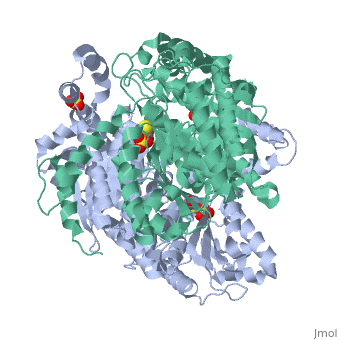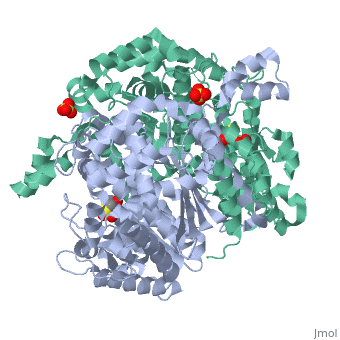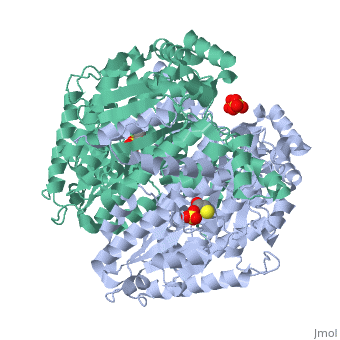
CRYSTAL STRUCTURE OF HUMAN PHOSPHOGLUCOSE ISOMERASE/NEUROLEUKIN/AUTOCRINE MOTILITY FACTOR/MATURATION FACTOR
Contents |
Overview
Phosphoglucose isomerase (PGI) is a multifunctional protein, which, inside the cell, functions as a housekeeping enzyme of glycolysis and gluconeogenesis and, outside the cell, exerts wholly unrelated cytokine properties. We have determined the structure of human PGI to a resolution of 1.6 A using X-ray crystallography. The structure is highly similar to other PGIs, especially the architecture of the active site. Fortuitous binding of a sulphate molecule from the crystallisation solution has facilitated an accurate description of the substrate phosphate-binding site. Comparison with both native and inhibitor-bound rabbit PGI structures shows that two loops move closer to the active site upon binding inhibitor. Interestingly, the human structure most closely resembles the inhibitor-bound structure, suggesting that binding of the phosphate moiety of the substrate may trigger this conformational change. We suggest a new mechanism for catalysis that uses Glu357 as the base catalyst for the isomerase reaction rather than His388 as proposed previously. The human PGI structure has also provided a detailed framework with which to map mutations associated with non-spherocytic haemolytic anaemia.
Disease
Known disease associated with this structure: Hemolytic anemia due to glucosephosphate isomerase deficiency OMIM:[172400], Hydrops fetalis, one form OMIM:[172400]
About this Structure
1IAT is a Single protein structure of sequence from Homo sapiens. Full crystallographic information is available from OCA.
Reference
The crystal structure of human phosphoglucose isomerase at 1.6 A resolution: implications for catalytic mechanism, cytokine activity and haemolytic anaemia., Read J, Pearce J, Li X, Muirhead H, Chirgwin J, Davies C, J Mol Biol. 2001 Jun 1;309(2):447-63. PMID:11371164
Page seeded by OCA on Sun Mar 30 21:17:07 2008