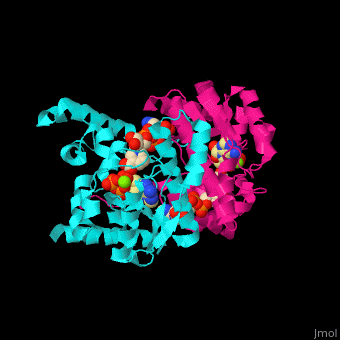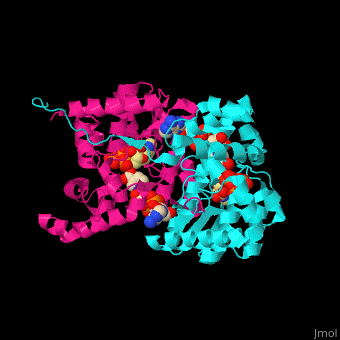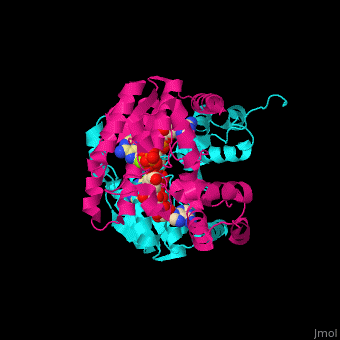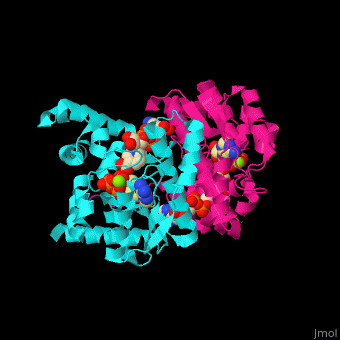This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
NAD synthase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | + | <StructureSection load='1xng' size='350' side='right' caption='NAD+ synthase complex with Mg+2 (green), ATP and deamino-NAD (PDB entry [[1xng]])' scene='54/541072/Cv/1'> | |
| - | <StructureSection load='1xng' size='350' side='right' caption='NAD+ synthase complex with Mg+2 (green), ATP | + | |
== Function == | == Function == | ||
Revision as of 11:13, 5 June 2016
| |||||||||||
3D Structures of NAD+ synthase
Updated on 05-June-2016
References
- ↑ Suda Y, Tachikawa H, Yokota A, Nakanishi H, Yamashita N, Miura Y, Takahashi N. Saccharomyces cerevisiae QNS1 codes for NAD(+) synthetase that is functionally conserved in mammals. Yeast. 2003 Aug;20(11):995-1005. PMID:12898714 doi:http://dx.doi.org/10.1002/yea.1008
- ↑ Kang GB, Kim YS, Im YJ, Rho SH, Lee JH, Eom SH. Crystal structure of NH3-dependent NAD+ synthetase from Helicobacter pylori. Proteins. 2005 Mar 1;58(4):985-8. PMID:15645437 doi:10.1002/prot.20377