This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
3igc
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| + | |||
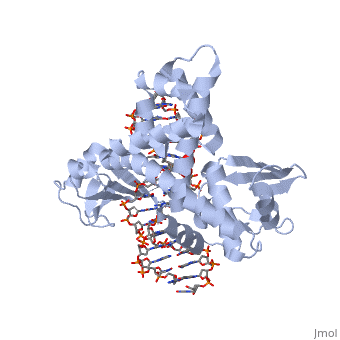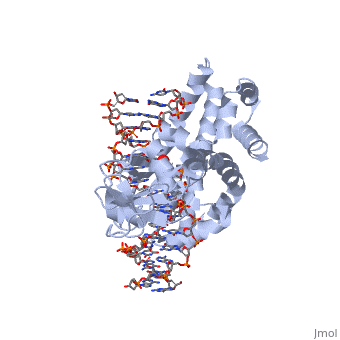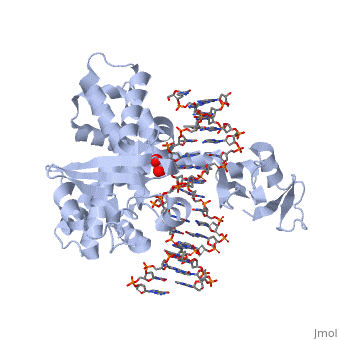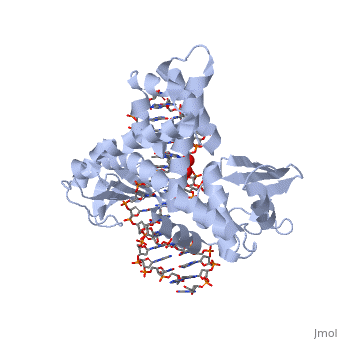
==Smallpox virus topoisomerase-DNA transition state== | ==Smallpox virus topoisomerase-DNA transition state== | ||
<StructureSection load='3igc' size='340' side='right' caption='[[3igc]], [[Resolution|resolution]] 2.10Å' scene=''> | <StructureSection load='3igc' size='340' side='right' caption='[[3igc]], [[Resolution|resolution]] 2.10Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[3igc]] is a 4 chain structure with sequence from [http://en.wikipedia.org/wiki/ | + | <table><tr><td colspan='2'>[[3igc]] is a 4 chain structure with sequence from [http://en.wikipedia.org/wiki/Varv Varv]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=3IGC OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=3IGC FirstGlance]. <br> |
</td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=VO4:VANADATE+ION'>VO4</scene></td></tr> | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=VO4:VANADATE+ION'>VO4</scene></td></tr> | ||
| - | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">H6R, I6R, TOP1 ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10255 | + | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">H6R, I6R, TOP1 ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10255 VARV])</td></tr> |
<tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/DNA_topoisomerase DNA topoisomerase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.99.1.2 5.99.1.2] </span></td></tr> | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/DNA_topoisomerase DNA topoisomerase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.99.1.2 5.99.1.2] </span></td></tr> | ||
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=3igc FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3igc OCA], [http://www.rcsb.org/pdb/explore.do?structureId=3igc RCSB], [http://www.ebi.ac.uk/pdbsum/3igc PDBsum]</span></td></tr> | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=3igc FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3igc OCA], [http://pdbe.org/3igc PDBe], [http://www.rcsb.org/pdb/explore.do?structureId=3igc RCSB], [http://www.ebi.ac.uk/pdbsum/3igc PDBsum], [http://prosat.h-its.org/prosat/prosatexe?pdbcode=3igc ProSAT]</span></td></tr> |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [[http://www.uniprot.org/uniprot/ | + | [[http://www.uniprot.org/uniprot/TOP1_VAR67 TOP1_VAR67]] Releases the supercoiling and torsional tension of DNA introduced during the DNA replication and transcription by transiently cleaving and rejoining one strand of the DNA duplex. Introduces a single-strand break via transesterification at the specific target site 5'-[CT]CCTTp site in duplex DNA. The scissile phosphodiester is attacked by the catalytic tyrosine of the enzyme, resulting in the formation of a DNA-(3'-phosphotyrosyl)-enzyme intermediate and the expulsion of a 5'-OH DNA strand. The free DNA strand then undergoes passage around the unbroken strand thus removing DNA supercoils. Finally, in the religation step, the DNA 5'-OH attacks the covalent intermediate to expel the active-site tyrosine and restore the DNA phosphodiester backbone (By similarity). |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 18: | Line 19: | ||
<text>to colour the structure by Evolutionary Conservation</text> | <text>to colour the structure by Evolutionary Conservation</text> | ||
</jmolCheckbox> | </jmolCheckbox> | ||
| - | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/ | + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=3igc ConSurf]. |
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
<div style="background-color:#fffaf0;"> | <div style="background-color:#fffaf0;"> | ||
| Line 28: | Line 29: | ||
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
</div> | </div> | ||
| + | <div class="pdbe-citations 3igc" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
| + | *[[Smallpox (Variola Virus) - Topoisomerase 1B|Smallpox (Variola Virus) - Topoisomerase 1B]] | ||
*[[Topoisomerase|Topoisomerase]] | *[[Topoisomerase|Topoisomerase]] | ||
*[[Variola Topoisomerase 1B|Variola Topoisomerase 1B]] | *[[Variola Topoisomerase 1B|Variola Topoisomerase 1B]] | ||
| Line 37: | Line 40: | ||
</StructureSection> | </StructureSection> | ||
[[Category: DNA topoisomerase]] | [[Category: DNA topoisomerase]] | ||
| - | [[Category: | + | [[Category: Varv]] |
[[Category: Bushman, F D]] | [[Category: Bushman, F D]] | ||
[[Category: Duyne, G D.Van]] | [[Category: Duyne, G D.Van]] | ||
Revision as of 23:29, 4 August 2016
Smallpox virus topoisomerase-DNA transition state
| |||||||||||