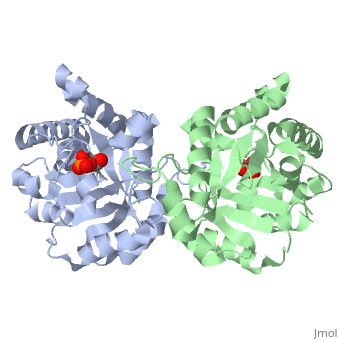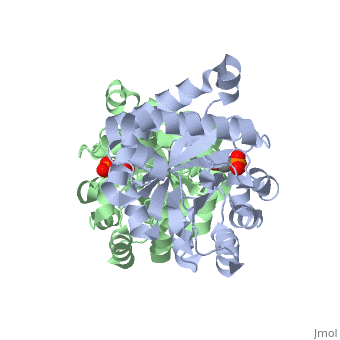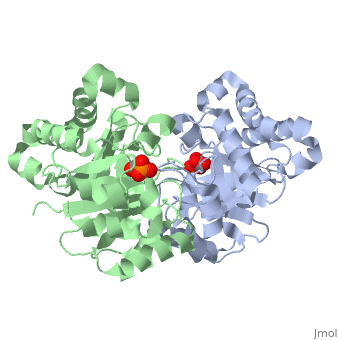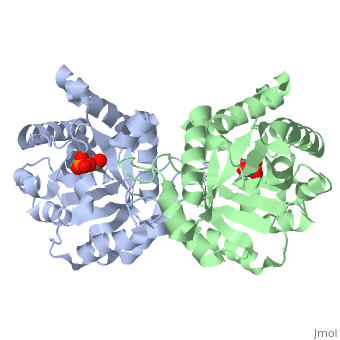This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2ypi
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| + | |||
==CRYSTALLOGRAPHIC ANALYSIS OF THE COMPLEX BETWEEN TRIOSEPHOSPHATE ISOMERASE AND 2-PHOSPHOGLYCOLATE AT 2.5-ANGSTROMS RESOLUTION. IMPLICATIONS FOR CATALYSIS== | ==CRYSTALLOGRAPHIC ANALYSIS OF THE COMPLEX BETWEEN TRIOSEPHOSPHATE ISOMERASE AND 2-PHOSPHOGLYCOLATE AT 2.5-ANGSTROMS RESOLUTION. IMPLICATIONS FOR CATALYSIS== | ||
<StructureSection load='2ypi' size='340' side='right' caption='[[2ypi]], [[Resolution|resolution]] 2.50Å' scene=''> | <StructureSection load='2ypi' size='340' side='right' caption='[[2ypi]], [[Resolution|resolution]] 2.50Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[2ypi]] is a 2 chain structure with sequence from [http://en.wikipedia.org/wiki/ | + | <table><tr><td colspan='2'>[[2ypi]] is a 2 chain structure with sequence from [http://en.wikipedia.org/wiki/Atcc_18824 Atcc 18824]. The February 2004 RCSB PDB [http://pdb.rcsb.org/pdb/static.do?p=education_discussion/molecule_of_the_month/index.html Molecule of the Month] feature on ''The Glycolytic Enzymes'' by David S. Goodsell is [http://dx.doi.org/10.2210/rcsb_pdb/mom_2004_2 10.2210/rcsb_pdb/mom_2004_2]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2YPI OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2YPI FirstGlance]. <br> |
</td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=PGA:2-PHOSPHOGLYCOLIC+ACID'>PGA</scene></td></tr> | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=PGA:2-PHOSPHOGLYCOLIC+ACID'>PGA</scene></td></tr> | ||
<tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/Triose-phosphate_isomerase Triose-phosphate isomerase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.3.1.1 5.3.1.1] </span></td></tr> | <tr id='activity'><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/Triose-phosphate_isomerase Triose-phosphate isomerase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=5.3.1.1 5.3.1.1] </span></td></tr> | ||
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2ypi FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2ypi OCA], [http://www.rcsb.org/pdb/explore.do?structureId=2ypi RCSB], [http://www.ebi.ac.uk/pdbsum/2ypi PDBsum]</span></td></tr> | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2ypi FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2ypi OCA], [http://pdbe.org/2ypi PDBe], [http://www.rcsb.org/pdb/explore.do?structureId=2ypi RCSB], [http://www.ebi.ac.uk/pdbsum/2ypi PDBsum], [http://prosat.h-its.org/prosat/prosatexe?pdbcode=2ypi ProSAT]</span></td></tr> |
</table> | </table> | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
| Line 15: | Line 16: | ||
<text>to colour the structure by Evolutionary Conservation</text> | <text>to colour the structure by Evolutionary Conservation</text> | ||
</jmolCheckbox> | </jmolCheckbox> | ||
| - | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/ | + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=2ypi ConSurf]. |
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
<div style="background-color:#fffaf0;"> | <div style="background-color:#fffaf0;"> | ||
| Line 25: | Line 26: | ||
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
</div> | </div> | ||
| + | <div class="pdbe-citations 2ypi" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
| Line 32: | Line 34: | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
| + | [[Category: Atcc 18824]] | ||
[[Category: RCSB PDB Molecule of the Month]] | [[Category: RCSB PDB Molecule of the Month]] | ||
| - | [[Category: Saccharomyces cerevisiae]] | ||
[[Category: The Glycolytic Enzymes]] | [[Category: The Glycolytic Enzymes]] | ||
[[Category: Triose-phosphate isomerase]] | [[Category: Triose-phosphate isomerase]] | ||
Revision as of 07:58, 11 August 2016
CRYSTALLOGRAPHIC ANALYSIS OF THE COMPLEX BETWEEN TRIOSEPHOSPHATE ISOMERASE AND 2-PHOSPHOGLYCOLATE AT 2.5-ANGSTROMS RESOLUTION. IMPLICATIONS FOR CATALYSIS
| |||||||||||