This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
SARS Coronavirus Main Proteinase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
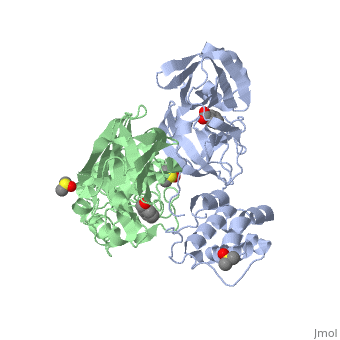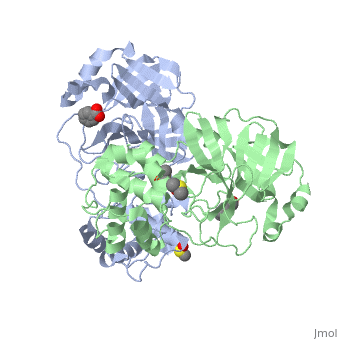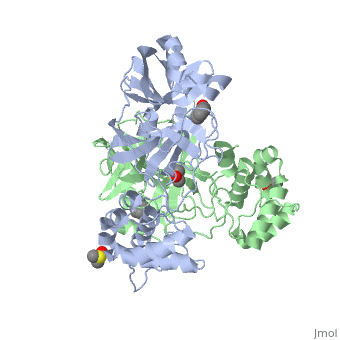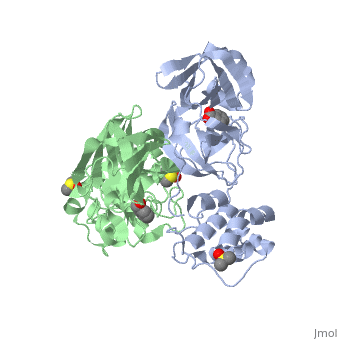
{{STRUCTURE_2vj1| PDB=2vj1 | SIZE=400| SCENE= |right|CAPTION=Human SARS coronavirus proteinase dimer complex with benzoic acid, dimethylamino benzoic acid and DMSO, [[2vj1]] }} | {{STRUCTURE_2vj1| PDB=2vj1 | SIZE=400| SCENE= |right|CAPTION=Human SARS coronavirus proteinase dimer complex with benzoic acid, dimethylamino benzoic acid and DMSO, [[2vj1]] }} | ||
| + | <StructureSection load='2vj1' size='350' side='right' caption='Human SARS coronavirus proteinase dimer complex with benzoic acid, dimethylamino benzoic acid and DMSO, [[2vj1]]' scene=''> | ||
==Introduction== | ==Introduction== | ||
Before the emergence of the SARS-CoV, no research efforts were being directed toward the discovery of antiviral drugs against coronaviruses. The high infectivity and mortality rate established SARS as a significant global threat for which no efficacious therapy was available. Since the epidemic, which began in 2003, new insights into the genomics and pathogenisis of the SARS-CoV revealed several novel potential anti-coronavirus targets. Several proteins encoded by the SARS-CoV could be considered as targets for therapeutic intervention. The focus of this article is the '''SARS-CoV main protease'''. | Before the emergence of the SARS-CoV, no research efforts were being directed toward the discovery of antiviral drugs against coronaviruses. The high infectivity and mortality rate established SARS as a significant global threat for which no efficacious therapy was available. Since the epidemic, which began in 2003, new insights into the genomics and pathogenisis of the SARS-CoV revealed several novel potential anti-coronavirus targets. Several proteins encoded by the SARS-CoV could be considered as targets for therapeutic intervention. The focus of this article is the '''SARS-CoV main protease'''. | ||
| Line 24: | Line 25: | ||
<scene name='SARS_Coronavirus_Main_Proteinase/Figure_5a/1'>Active-site reacted with 1-(dimethylaminobenzoyloxy)-benzotriazole</scene> | <scene name='SARS_Coronavirus_Main_Proteinase/Figure_5a/1'>Active-site reacted with 1-(dimethylaminobenzoyloxy)-benzotriazole</scene> | ||
| - | + | </StructureSection> | |
==3D structures of virus proteinase== | ==3D structures of virus proteinase== | ||
| Line 32: | Line 33: | ||
*Coronavirus protease | *Coronavirus protease | ||
| - | **[[1q2w]], [[3iwm]], [[3ebn]], [[3d23]], [[ | + | **[[1q2w]], [[3iwm]], [[3ebn]], [[3d23]], [[5b6o]], [[2gt7]], [[1z1i]], [[2a5a]], [[2c3s]], [[2bx3]], [[2bx4]], [[1uj1]], [[1uk2]], [[1uk3]], [[2duc]], [[2yna]] – 3CLP - SARS coronavirus<br /> |
**[[2liz]] – 3CLP C terminal – NMR<br /> | **[[2liz]] – 3CLP C terminal – NMR<br /> | ||
**[[2amd]], [[2amq]], [[2amp]], [[1wof]], [[2ynb]], [[3atw]], [[3avz]] – 3CLP + inhibitor <br /> | **[[2amd]], [[2amq]], [[2amp]], [[1wof]], [[2ynb]], [[3atw]], [[3avz]] – 3CLP + inhibitor <br /> | ||
Revision as of 11:07, 24 November 2016
| |||||||||||
3D structures of virus proteinase
Updated on 24-November-2016
References
- ↑ Murray, Patrick R., Ken S. Rosenthal, and Michael A. Pfaller. "Coronaviruses and Noroviruses." Medical Microbiology. 6th ed. Philadelphia: Mosby/Elsevier, 2009. 565-68. Print.
- ↑ Ziebuhr, J., Herold,J., and Siddell, S.G. (1995). Characterization of a human coronavirus (strain 229E) 3C-like proteinase activity. J. Virol. 69, 4331-4338.
- ↑ Ksiazek TG, Erdman D, Goldsmith CS, et al. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med 2003;348:1953-1966
- ↑ "Summary of probable SARS cases with onset of illness from 1 November 2002 to 31 July 2003". WHO. Retrieved 2008-10-31.
- ↑ Li W, Shi Z, Yu M, et al. (2005). "Bats are natural reservoirs of SARS-like coronaviruses". Science (journal) 310 (5748): 676–9. doi:10.1126/science.1118391. PMID 16195424.
- ↑ Ziebuhr, J., Snijder, E.J., and Gorbaleya, A.E. (2000). Virus-encoded proteinases and proteolytic processing in Nidovirales. J. Gen. Virol. 81, 853-879.
- ↑ Anad, K., Ziebhr, J., Wadhani, P., Mesters, J.R., and Hilgenfeld, R. (2003). Coronavirus main protease (3CLPro) structure: basis for design of anti-SARS drugs. Science300, 1763-1767.
- ↑ Anad, K., Ziebhr, J., Wadhani, P., Mesters, J.R., and Hilgenfeld, R. (2003). Coronavirus main protease (3CLPro) structure: basis for design of anti-SARS drugs. Science300, 1763-1767.
- ↑ Verschueren, Koen H.G., Ksenia Pumpor, Stefan Anemüller, Shuai Chen, Jeroen R. Mesters, and Rolf Hilgenfeld. "A Structural View of the Inactivation of the SARS Coronavirus Main Proteinase by Benzotriazole Esters." Chemistry & Biology 15.6 (2008): 597-606. Web. Oct. 2010.
Proteopedia Page Contributors and Editors (what is this?)
Sarra Borhanian, Michal Harel, David Canner, Ann Taylor, Alexander Berchansky