This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Reverse transcriptase
From Proteopedia
(Difference between revisions)
| Line 18: | Line 18: | ||
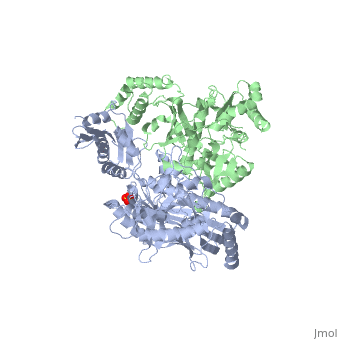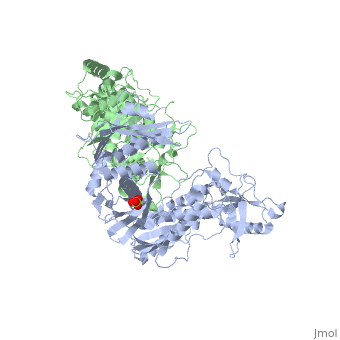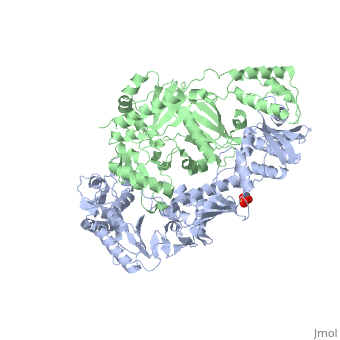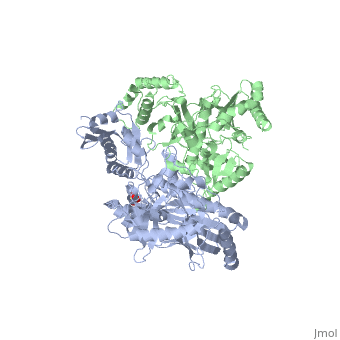
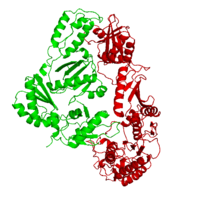
This ''hand-like'' <scene name='Reverse_transcriptase/Chains/2'>heterodimer</scene> protein has an usual length of 1000 residues (560 in Chain A and 440 for B), a third of them involved in alpha helices and almost a quarter involved in beta sheets, showing α+β <scene name='Reverse_transcriptase/Secondary/2'>secondary structure</scene> domains. <scene name='Reverse_transcriptase/Chaina/2'>Chain A</scene> has an usual weight of 66KDa whereas <scene name='Reverse_transcriptase/Chainb/2'>Chain B</scene> is around 51KDa. These monomers are derived from the same gene, but p51 lacks the amino acids of one active site and has a different tertiary structure conformation compared to p66. Because of this, p51 is enzymatically inactive<ref>PMID: 1377403</ref> | This ''hand-like'' <scene name='Reverse_transcriptase/Chains/2'>heterodimer</scene> protein has an usual length of 1000 residues (560 in Chain A and 440 for B), a third of them involved in alpha helices and almost a quarter involved in beta sheets, showing α+β <scene name='Reverse_transcriptase/Secondary/2'>secondary structure</scene> domains. <scene name='Reverse_transcriptase/Chaina/2'>Chain A</scene> has an usual weight of 66KDa whereas <scene name='Reverse_transcriptase/Chainb/2'>Chain B</scene> is around 51KDa. These monomers are derived from the same gene, but p51 lacks the amino acids of one active site and has a different tertiary structure conformation compared to p66. Because of this, p51 is enzymatically inactive<ref>PMID: 1377403</ref> | ||
| - | There are five distinct structures within the p66 subchain that are used to describe the functions of RT: the fingers (residues 1–85 and 118–155), the palm (residues 86–117 and 156–236), the thumb (residues 237–318), the connection (319–426), and the RNase H (residues 427-end). The palm contains the main active site (residues 110, 185-186) | + | There are five distinct structures within the p66 subchain that are used to describe the functions of RT: the fingers (residues 1–85 and 118–155), the palm (residues 86–117 and 156–236), the thumb (residues 237–318), the connection (319–426), and the RNase H (residues 427-end). The palm contains the main active site (residues 110, 185-186)<ref> PMID:2881421</ref> |
{{Clear}} | {{Clear}} | ||
Revision as of 13:53, 27 September 2017
| |||||||||||
3D Structures of Reverse transcriptase
Updated on 27-September-2017
See Also
- Reverse Transcriptase at Wikipedia
- Molecule of the Month (09/2002) at RCSB Protein Data Bank
- List of Reverse Transcriptase articles at Proteopedia and at RCSB Protein Data Bank
- Model of Reverse Transcriptase as one of the CBI Molecules on the Molecular Playground
- See Transcription for additional Proteopedia articles on the subject.
- For additional information, see: Human Immunodeficiency Virus
- For additional information, see: Transcription and RNA Processing
References
- ↑ Kohlstaedt LA, Wang J, Friedman JM, Rice PA, Steitz TA. Crystal structure at 3.5 A resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science. 1992 Jun 26;256(5065):1783-90. PMID:1377403 doi:[http://dx.doi.org/10.1126/science.1377403 http://dx.doi.org/10.1126/science.1377403
- ↑ Kondo H, Terada H, Iseki T, Iwasa S, Okuda K, Kanazawa S, Okuda K. [Effects of cobalamin analogues produced by reaction between OHB12 and ascorbic acid upon absorption, plasma transport, organ distribution and B12 dependent enzymes]. Nihon Ketsueki Gakkai Zasshi. 1986 Nov;49(7):1338-46. PMID:2881421
- ↑ ConSurf: Using Evolutionary Data to Raise Testable Hypotheses about Protein Function DOI: 10.1002/ijch.201200096
- ↑ Abbondanzieri EA, Bokinsky G, Rausch JW, Zhang JX, Le Grice SF, Zhuang X. Dynamic binding orientations direct activity of HIV reverse transcriptase. Nature. 2008 May 8;453(7192):184-9. PMID:18464735 doi:10.1038/nature06941
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Daniel Moyano-Marino, Joel L. Sussman, Alexander Berchansky, David Canner, Amol Kapoor, Jaime Prilusky, Brian Foley, Lynmarie K Thompson, Eric Martz