Nuclear polyadenylated RNA-binding protein
From Proteopedia
| Line 10: | Line 10: | ||
[[Image:Conserved W168 in Hrp1.png|300 px|right|thumb|Figure 2: Sequence logo for residues 167-169 of Hrp1. The logo displays the frequency of residues occuring at specific positions within Hrp1. W168 is always conserved in Hrp1 and RRMs of similar proteins.]] | [[Image:Conserved W168 in Hrp1.png|300 px|right|thumb|Figure 2: Sequence logo for residues 167-169 of Hrp1. The logo displays the frequency of residues occuring at specific positions within Hrp1. W168 is always conserved in Hrp1 and RRMs of similar proteins.]] | ||
| + | |||
==RBD-RBD Interactions and the Linker Region== | ==RBD-RBD Interactions and the Linker Region== | ||
As mentioned above, Hrp1 is composed of two RBDs. The RBDs are connected by a linker region which also contains an crucial residue for RNA binding. Ile234 holds Ade6 stacked in place with Phe162 <scene name='78/781945/Linker_rna/1'>via van der Waals contacts</scene>. Experimental evidence from the NMR data <ref name="GM3H"/> suggests that the two RBDs act independently until binding the PEE. Binding the PEE causes the linker region to adopt a short helical structure to rigidly hold the <scene name='78/781945/Protein_domains/2'>RBDs in place relative to each other</scene>. Aside from the linker helix, the only interaction between the RBDs is due to <scene name='78/781945/Interaction_between_domains/5'>a single salt bridge</scene> between Lys231 and Asp271. | As mentioned above, Hrp1 is composed of two RBDs. The RBDs are connected by a linker region which also contains an crucial residue for RNA binding. Ile234 holds Ade6 stacked in place with Phe162 <scene name='78/781945/Linker_rna/1'>via van der Waals contacts</scene>. Experimental evidence from the NMR data <ref name="GM3H"/> suggests that the two RBDs act independently until binding the PEE. Binding the PEE causes the linker region to adopt a short helical structure to rigidly hold the <scene name='78/781945/Protein_domains/2'>RBDs in place relative to each other</scene>. Aside from the linker helix, the only interaction between the RBDs is due to <scene name='78/781945/Interaction_between_domains/5'>a single salt bridge</scene> between Lys231 and Asp271. | ||
Revision as of 03:32, 29 March 2018
Contents |
Introduction
|
Hrp1 is a polyadenylation factor found in Saccharomyces cervisiae (yeast) [1]. This protein recognizes and binds to an RNA sequence in the 3'UTR upstream from the cleavage site called the polyadenylation enhancement element (PEE) [1]. Upon binding to the RNA, Hrp1 helps recruit additional proteins necessary for the cleavage and polyadenylation of the RNA molecule [1]. The structure of the Hrp1-PEE complex reveals the mechanism by which Hrp1 is able to recognize and bind to its specific RNA sequence at the atomic level [1].
Structure
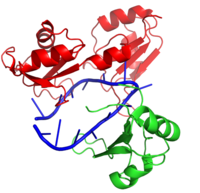
Hrp1 is a single strand RNA-binding protein composed of two RNP-type RNA-binding domains (RBDs) arranged in tandem with a typical ßαßßαß architecture [1]. The two RBDs have similar topolgies, both containing a central antiparallel four-stranded with two α-helices running across one face [1]. The two RBDs associate to form a deep and positively charged , which constitutes the binding site for the RNA molecule [1].
Hrp1-RNA Interactions
The interface between Hrp1 and its target RNA sequence is dominated by interactions between key aromatic residues and RNA bases [1]. Only six RNA bases, an repeat, act as the PEE and form specific contacts with Hrp1 [1]. Hydrophilic residues of Hrp1 provide base specificity through hydrogen bonding [1]. Most of the key residues that interact with the RNA can be found in the ß-sheet region of Hrp1; however, loops and the interdomain linker are also essential for Hrp1-RNA recognition [1]. Perhaps the most important Hrp1-RNA interaction is the (a conserved residue) [1]. In this case, Trp168 stacks on Ade4 and forms crucial base-specific hydrogen bonds [1]. It is also worth noting that a second Hrp1 residue is critical to holding Ade4 in place, , which interacts via hydrogen bond with the N1 of Ade4 [1]. A third contributor, , also stacks with Ura7 to aid in RNA recognition and binding [1].
RBD-RBD Interactions and the Linker Region
As mentioned above, Hrp1 is composed of two RBDs. The RBDs are connected by a linker region which also contains an crucial residue for RNA binding. Ile234 holds Ade6 stacked in place with Phe162 . Experimental evidence from the NMR data [1] suggests that the two RBDs act independently until binding the PEE. Binding the PEE causes the linker region to adopt a short helical structure to rigidly hold the . Aside from the linker helix, the only interaction between the RBDs is due to between Lys231 and Asp271.
Relationship to other proteins
The RNP-type RBD is found in many proteins involved in post-transcriptional pre-mRNA processing (5' end capping, splicing, 3' end polyadenylation, and transport from the nucleus)[2]. The RBD of Hrp1 enables the protein to bind RNA sequences that differ in both length and content from the RBDs of other mRNA processing proteins such as sex lethal, PAPB, and HuD [1].
Interaction with RNA15
RNA15 is another RNA-binding protein with a single N-terminal RNA recognition motif (RRM) [3]. RNA15 recognizes an A-rich positioning element (PE) downstream from the PEE but upstream from the 3' cleavage site [3]. The recognition of the PE by RNA15 is crucial for precise cleavage of the RNA molecule. Hrp1 and RNA15 are held together by a separate protein, RNA14 [3]. These proteins act together to anchor the polyadenylation and cleavage protein machinery relative to the cleavage site for precise 3'-end processing [3].
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 Perez-Canadillas JM. Grabbing the message: structural basis of mRNA 3'UTR recognition by Hrp1. EMBO J. 2006 Jul 12;25(13):3167-78. Epub 2006 Jun 22. PMID:16794580
- ↑ Clery A, Blatter M, Allain FH. RNA recognition motifs: boring? Not quite. Curr Opin Struct Biol. 2008 Jun;18(3):290-8. doi: 10.1016/j.sbi.2008.04.002. PMID:18515081 doi:http://dx.doi.org/10.1016/j.sbi.2008.04.002
- ↑ 3.0 3.1 3.2 3.3 Leeper TC, Qu X, Lu C, Moore C, Varani G. Novel protein-protein contacts facilitate mRNA 3'-processing signal recognition by Rna15 and Hrp1. J Mol Biol. 2010 Aug 20;401(3):334-49. Epub 2010 Jun 19. PMID:20600122 doi:10.1016/j.jmb.2010.06.032
Proteopedia Page Contributors and Editors (what is this?)
Cory A. Wuerch, Matthew Douglas Moore, Savannah Davis, Michal Harel, Jaime Prilusky