|
Introduction
Purine-rich element binding protein alpha (Purα) is a transcription factor with a molecular weight of ~35 kDa encoded by the PURA gene. It possesses ATP-independent dsDNA unwinding activity, and is known to bind sequence-specific purine-rich regions of ssDNA and ssRNA, recognizing GGN motifs. Purα is a member of the PUR family of proteins, which includes Purβ and two isoforms of Purγ. In its functional dimeric form Purα is known to repress expression of smooth muscle alpha actin (SMαA) encoded by the Acta2 gene. It is also known to be involved in DNA replication and cell cycle regulation as well as mRNA translation. It plays a crucial role in nervous system development, and mutations in Purα have been implicated in two neurological diseases: PURA syndrome and Fragile X-associated Tremor/Ataxia Syndrome (FXTAS) (see Disease section).[1]
Structure
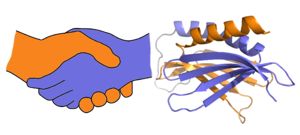 A PUR domain is analogous to a left-handed handshake. PUR repeat I-II represented from 5fgp. Purα functions as a dimer composed of two intramolecular domains and one intermolecular domain. The Purα monomer contains three semi-conserved repeated amino acid sequences, named in order from N->C: PUR repeats I, II, and III. These repeats fold to form two domains: associating to form the I-II domain or “intramolecular domain”, while facilitates dimerization through association with a repeat III from a second Purα monomer or repeat III of Purβ. Each PUR repeat is connected by flexible linker regions. Each PUR repeat contains a beta-sheet composed of four beta-strands, followed by a single alpha-helix. While Purα is not yet officially classified by SCOP or CATH, its structure is that of an α+β protein.
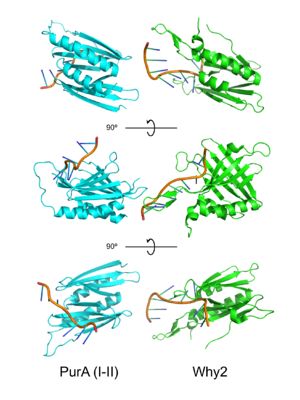 PUR domains are structurally similar to the fold in Whirly proteins. Left: PUR repeat I-II ( 5fgp), right: WHY2 ( 3n1k). The domains of Purα have been described as "Whirly-like" folds because of their structural similarity to the DNA-binding Whirly class of proteins found in plants.[2] Whirlys are also ssDNA binding proteins, however unlike Purα they are not sequence-specific.
The 2016 X-ray crystal structure of Purα repeat I-II (5fgp) shows DNA bound between repeats I and II. Residues involved in protein-DNA conjugation are: (repeat I) Q52, S53 and K54; (repeat II) K138, N140, R142, and F145. K54 (repeat I) and K138 (repeat II) interact with the phosphate backbone, while Q52 (repeat I), S53 (repeat I), K54 (repeat I), R142 (repeat II), and N140 (repeat II) hydrogen bond directly with the DNA bases. Interestingly all bases appear to be stabilized by stacking interactions within the DNA strand except G4 which stacks directly with F145 (repeat II) of the protein. This phenylalanine-guanine stacking interaction disrupts base-stacking within the DNA resulting in the following cytosine being flipped out, introducing a strong kink in the DNA strand. This base-stacking interruption was shown to be responsible for Purα's DNA-unwinding activity.[3]
Function
High-affinity nucleic acid-binding function is dependent on Purα dimerization. Purα forms homodimers in addition to heterodimers with Purβ. PurA is known to repress various genes including
, Y57 (repeat I) and F145 (repeat II) have been implicated in the DNA unwinding activity of PurA.[4]
Development
Purα expression is highest during fetal development, specifically in the brain, spinal cord, and genitalia. In humans and mice Purα expression decreases in adulthood, and is restricted primarily to the brain and testes.
Tissue Specificity
While Purα is a ubiquitously expressed protein in all cell types, it is generally found at low levels in most tissues. However, in adult humans and mice Purα expression is highest in the brain and spinal cord, and to a lesser extent in the testes. Purα has been shown to be upregulated in cardiac myocyte gap junctions following heart transplant in mice, colocalizing with SMαA.[5]
Disease
Mutations in the PURA gene resulting in haploinsufficiency of Purα are known to cause the neurological disease PURA syndrome. PURA syndrome appears early in development, , with patients exhibiting severe developmental delay, seizures, feeding difficulty, of severe intellectual disabilities, movements, vision, hypotonia, premature telarche, . This disease has no cure, and life expectancy is .[6]. Purα knock-out mice exhibit similar neurological symptoms such as severe tremor developing at about postnatal week two, and feeding difficulties. These mice die after approximately one month. Heterozygous mice display less severe symptoms including seizures upon handling. [7]
Fragile X-associated Tremor/Ataxia Syndrome (FXTAS) is another disease associated with abnormal activity of Purα. Normally between __ and __ copies of __ , however in greater than __ copies can result in ___ causing delayed-onset neurological problems. FXTAS generally develops in middle age, and is characterized by tremor, ataxia, and ___.
JSmol in Proteopedia [8]
References
- ↑ Weber J, Bao H, Hartlmuller C, Wang Z, Windhager A, Janowski R, Madl T, Jin P, Niessing D. Structural basis of nucleic-acid recognition and double-strand unwinding by the essential neuronal protein Pur-alpha. Elife. 2016 Jan 8;5. pii: e11297. doi: 10.7554/eLife.11297. PMID:26744780 doi:http://dx.doi.org/10.7554/eLife.11297
- ↑ Graebsch A, Roche S, Niessing D. X-ray structure of Pur-alpha reveals a Whirly-like fold and an unusual nucleic-acid binding surface. Proc Natl Acad Sci U S A. 2009 Nov 3;106(44):18521-6. Epub 2009 Oct 21. PMID:19846792
- ↑ Weber J, Bao H, Hartlmuller C, Wang Z, Windhager A, Janowski R, Madl T, Jin P, Niessing D. Structural basis of nucleic-acid recognition and double-strand unwinding by the essential neuronal protein Pur-alpha. Elife. 2016 Jan 8;5. pii: e11297. doi: 10.7554/eLife.11297. PMID:26744780 doi:http://dx.doi.org/10.7554/eLife.11297
- ↑ Weber J, Bao H, Hartlmuller C, Wang Z, Windhager A, Janowski R, Madl T, Jin P, Niessing D. Structural basis of nucleic-acid recognition and double-strand unwinding by the essential neuronal protein Pur-alpha. Elife. 2016 Jan 8;5. pii: e11297. doi: 10.7554/eLife.11297. PMID:26744780 doi:http://dx.doi.org/10.7554/eLife.11297
- ↑ Zhang A, David JJ, Subramanian SV, Liu X, Fuerst MD, Zhao X, Leier CV, Orosz CG, Kelm RJ Jr, Strauch AR. Serum response factor neutralizes Pur alpha- and Pur beta-mediated repression of the fetal vascular smooth muscle alpha-actin gene in stressed adult cardiomyocytes. Am J Physiol Cell Physiol. 2008 Mar;294(3):C702-14. doi:, 10.1152/ajpcell.00173.2007. Epub 2008 Jan 9. PMID:18344281 doi:http://dx.doi.org/10.1152/ajpcell.00173.2007
- ↑ Reijnders MRF, Janowski R, Alvi M, Self JE, van Essen TJ, Vreeburg M, Rouhl RPW, Stevens SJC, Stegmann APA, Schieving J, Pfundt R, van Dijk K, Smeets E, Stumpel CTRM, Bok LA, Cobben JM, Engelen M, Mansour S, Whiteford M, Chandler KE, Douzgou S, Cooper NS, Tan EC, Foo R, Lai AHM, Rankin J, Green A, Lonnqvist T, Isohanni P, Williams S, Ruhoy I, Carvalho KS, Dowling JJ, Lev DL, Sterbova K, Lassuthova P, Neupauerova J, Waugh JL, Keros S, Clayton-Smith J, Smithson SF, Brunner HG, van Hoeckel C, Anderson M, Clowes VE, Siu VM, Ddd Study T, Selber P, Leventer RJ, Nellaker C, Niessing D, Hunt D, Baralle D. PURA syndrome: clinical delineation and genotype-phenotype study in 32 individuals with review of published literature. J Med Genet. 2018 Feb;55(2):104-113. doi: 10.1136/jmedgenet-2017-104946. Epub, 2017 Nov 2. PMID:29097605 doi:http://dx.doi.org/10.1136/jmedgenet-2017-104946
- ↑ Hunt D, Leventer RJ, Simons C, Taft R, Swoboda KJ, Gawne-Cain M, Magee AC, Turnpenny PD, Baralle D. Whole exome sequencing in family trios reveals de novo mutations in PURA as a cause of severe neurodevelopmental delay and learning disability. J Med Genet. 2014 Dec;51(12):806-13. doi: 10.1136/jmedgenet-2014-102798. Epub, 2014 Oct 23. PMID:25342064 doi:http://dx.doi.org/10.1136/jmedgenet-2014-102798
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
|


