This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Jennifer Taylor/Sandbox 3
From Proteopedia
(Difference between revisions)
| Line 18: | Line 18: | ||
== Methods == | == Methods == | ||
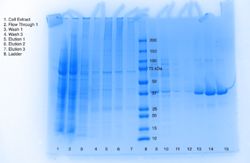
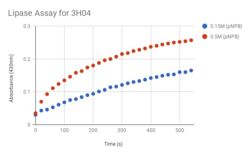
| - | We performed a Lipase Assay to determine what kind of esterase our protein is, because our target homolog (1TAH) is a lipase. The literature about 1TAH had no useful information about assays, but we used an assay from a paper regarding a different lipase. We knew that our protein was a hydrolase, and we believed it to be an esterase, and with this lipase assay we were able to deduce that our protein was an esterase and a lipase. | + | We performed a Lipase Assay to determine what kind of esterase our protein is, because our target homolog (1TAH) is a lipase. The literature about 1TAH had no useful information about assays, but we used an assay from a paper regarding a different lipase. We knew that our protein was a hydrolase, and we believed it to be an esterase, and with this lipase assay we were able to deduce that our protein was an esterase and a lipase. To determine if 3H04 functions as a lipase on the p-Nitrophenyl Butyrate (pNPB), a colorimetric assay using a spectrophotometry protocol. After mixing pNPB in 2-propanol, add 50 mM Tris-HCl buffer (pH 8.0, containing 0.4% w/v Triton XI00 and 0.1% w/v arabic gum) to the mixture. To initiate reaction, add pNPB solution and measure absorbance using a UV spectrophotometer at 410 nm for 5-10 min, recording absorbance values at 20 second intervals. |
== Results == | == Results == | ||
| Line 30: | Line 30: | ||
== Conclusion == | == Conclusion == | ||
| + | We've concluded that our protein is a lipase, and because we expressed our protein through E.coli, we believe that 3H04 functions in the lower intestine of warm blooded organisms. | ||
This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | This is a sample scene created with SAT to <scene name="/12/3456/Sample/1">color</scene> by Group, and another to make <scene name="/12/3456/Sample/2">a transparent representation</scene> of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes. | ||
Revision as of 18:13, 20 May 2018
3H04 Test Page
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644