This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Monooxygenase
From Proteopedia
(Difference between revisions)
| Line 57: | Line 57: | ||
**[[4j2w]], [[4j31]], [[4j33]], [[4j34]] – yKMO + FAD – yeast<br /> | **[[4j2w]], [[4j31]], [[4j33]], [[4j34]] – yKMO + FAD – yeast<br /> | ||
| - | **[[4j36]] – yKMO + FAD + inhibitor<br /> | + | **[[4j36]], [[5x64]] – yKMO + FAD + inhibitor<br /> |
| - | **[[5na5]] – PfKMO (mutant) + FAD – ''Pseudomonas fluorescens''<br /> | + | **[[5x6q]] – yKMO (mutant) + FAD + inhibitor<br /> |
| + | **[[5x68]] – hKMO + FAD - human<br /> | ||
| + | **[[5na5]], [[5x6p]] – PfKMO (mutant) + FAD – ''Pseudomonas fluorescens''<br /> | ||
| + | **[[5y7a]], [[5y77]] – PfKMO + FAD + kynurenine <br /> | ||
**[[5nak]] – PfKMO (mutant) + FAD + kynurenine <br /> | **[[5nak]] – PfKMO (mutant) + FAD + kynurenine <br /> | ||
| + | **[[5y66]] – PfKMO + FAD + kynurenine + inhibitor<br /> | ||
**[[5nah]], [[5nag]], [[5nae]], [[5nab]], [[5n7t]], [[5mzk]], [[5mzi]], [[5mzc]], [[5fn0]] – PfKMO (mutant) + FAD + inhibitor<br /> | **[[5nah]], [[5nag]], [[5nae]], [[5nab]], [[5n7t]], [[5mzk]], [[5mzi]], [[5mzc]], [[5fn0]] – PfKMO (mutant) + FAD + inhibitor<br /> | ||
| Line 69: | Line 73: | ||
*'''Phenylalanine 2-monooxygenase''' | *'''Phenylalanine 2-monooxygenase''' | ||
| - | **[[5pah]], [[6pah]] | + | **[[5pah]], [[6pah]] – hFMO catalytic domain + inhibitor <br /> |
*'''ActVA-Orf6 monooxygenase (AOMO)''' | *'''ActVA-Orf6 monooxygenase (AOMO)''' | ||
| Line 98: | Line 102: | ||
**[[2xvh]], [[2xlu]], [[2xlt]] – MaMO + FAD + NADP derivative<br /> | **[[2xvh]], [[2xlu]], [[2xlt]] – MaMO + FAD + NADP derivative<br /> | ||
**[[2xls]], [[2xlr]], [[2xlp]], [[2vq7]] – MaMO (mutant) + FAD + NADP <br /> | **[[2xls]], [[2xlr]], [[2xlp]], [[2vq7]] – MaMO (mutant) + FAD + NADP <br /> | ||
| - | + | **[[5nmw]] – ZvMO + FAD – ''Zonocerus variegatus''<br /> | |
| - | + | **[[5nmx]] – ZvMO + FAD + NADP <br /> | |
| - | + | ||
| - | **[[ | + | |
| - | **[[ | + | |
*2,4,6-trichlorophenol 4-monooxygenase | *2,4,6-trichlorophenol 4-monooxygenase | ||
| Line 153: | Line 154: | ||
**[[4q4k]] – PaMO + FMN <br /> | **[[4q4k]] – PaMO + FMN <br /> | ||
**[[5lsm]] – MO + FMN – ''Shewanella oneidensis''<br /> | **[[5lsm]] – MO + FMN – ''Shewanella oneidensis''<br /> | ||
| + | **[[6bka]] – MO + FMN – ''Cyberlyndnera mrakii''<br /> | ||
*2-hydroxbiphenyl 3-monooxygenase | *2-hydroxbiphenyl 3-monooxygenase | ||
| Line 192: | Line 194: | ||
**[[5cqf]] – MO – ''Pseudomonas syringae''<br /> | **[[5cqf]] – MO – ''Pseudomonas syringae''<br /> | ||
| - | **[[4d7e]] – | + | **[[4d7e]] – NfMO (mutant) + FAD – ''Nocardia farcinica''<br /> |
| + | **[[5o8p]] – EaMO + FAD – ''Erwinia amylovora''<br /> | ||
| + | **[[5o8r]] – EaMO + FAD + NADP <br /> | ||
*EDTA monooxygenase | *EDTA monooxygenase | ||
**[[5dqp]] – MO – ''Chelativorans''<br /> | **[[5dqp]] – MO – ''Chelativorans''<br /> | ||
| + | |||
| + | *Rifampicin monooxygenase or Pentachlorophenol 4-monooxygenase | ||
| + | |||
| + | **[[5vqb]] – SvMO + FAD – ''Sterptomyces venezuelae''<br /> | ||
| + | **[[6brd]] – SvMO (mutant) + FAD + rifampicin <br /> | ||
| + | **[[5kow]] – NfMO + FAD <br /> | ||
| + | **[[5kox]] – NfMO + FAD + rifampicin <br /> | ||
*3,6-diketocamphane 1,6 monooxygenase | *3,6-diketocamphane 1,6 monooxygenase | ||
| Line 212: | Line 223: | ||
**[[4mai]] – moMO + Cu – mold<br /> | **[[4mai]] – moMO + Cu – mold<br /> | ||
**[[4mah]] – moMO + Zn<br /> | **[[4mah]] – moMO + Zn<br /> | ||
| + | **[[5no7]] – MO – ''Pycnoporus cinnabarinus''<br /> | ||
*Peptidyl-glycine alpha-amidating monooxygenase | *Peptidyl-glycine alpha-amidating monooxygenase | ||
| + | **[[5mw0]], [[5wkw]] – rMO <br /> | ||
**[[3mib]], [[3mic]], [[3mid]], [[3mie]], [[3mif]], [[3mig]], [[3mih]], [[3mlj]], [[3mlk]], [[3mll]], [[1opm]] – rMO + Cu + Ni <br /> | **[[3mib]], [[3mic]], [[3mid]], [[3mie]], [[3mif]], [[3mig]], [[3mih]], [[3mlj]], [[3mlk]], [[3mll]], [[1opm]] – rMO + Cu + Ni <br /> | ||
**[[1sdw]] – rMO + Cu + Ni + O2 + threonine derivative<br /> | **[[1sdw]] – rMO + Cu + Ni + O2 + threonine derivative<br /> | ||
**[[1yjl]] – rMO (mutant)<br /> | **[[1yjl]] – rMO (mutant)<br /> | ||
**[[1yjk]], [[1yip]], [[1phm]] – rMO + Cu<br /> | **[[1yjk]], [[1yip]], [[1phm]] – rMO + Cu<br /> | ||
| - | **[[1yi9]] – rMO (mutant) + Cu<br /> | + | **[[1yi9]], [[6ay0]], [[6an3]], [[6amp]], [[6alv]], [[6ala]] – rMO (mutant) + Cu<br /> |
| + | **[[6ao6]], [[5wja]] – rMO (mutant) + Cu + Ni<br /> | ||
**[[3fw0]] – rMO + Hg<br /> | **[[3fw0]] – rMO + Hg<br /> | ||
**[[3fvz]] – rMO + Zn + Fe<br /> | **[[3fvz]] – rMO + Zn + Fe<br /> | ||
Revision as of 08:50, 12 August 2018
| |||||||||||
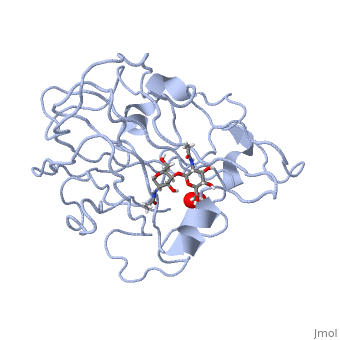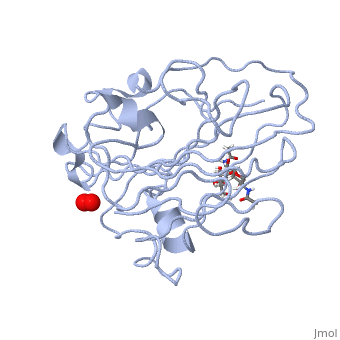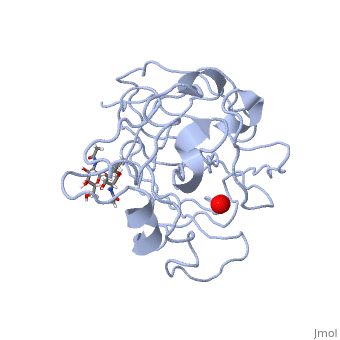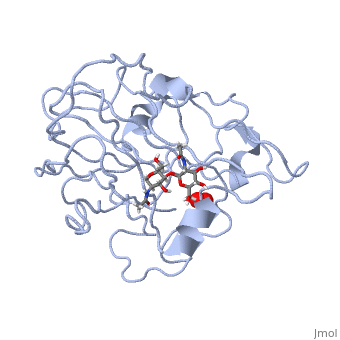
3D structures of monooxygenase
Updated on 12-August-2018
Methane monooxygenase See Methane monooxygenase
Camphor 5-monooxygenase See Cytochrome P450
Luciferin 4-monooxygenase and Alkanal monooxygenase See Luciferase
References
- ↑ Rudzka K, Moreno DM, Eipper B, Mains R, Estrin DA, Amzel LM. Coordination of peroxide to the Cu(M) center of peptidylglycine alpha-hydroxylating monooxygenase (PHM): structural and computational study. J Biol Inorg Chem. 2012 Dec 18. PMID:23247335 doi:10.1007/s00775-012-0967-z
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Joel L. Sussman, Alexander Berchansky, Jaime Prilusky