This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Poly(A) RNA polymerase
From Proteopedia
(Difference between revisions)
| Line 2: | Line 2: | ||
== Function == | == Function == | ||
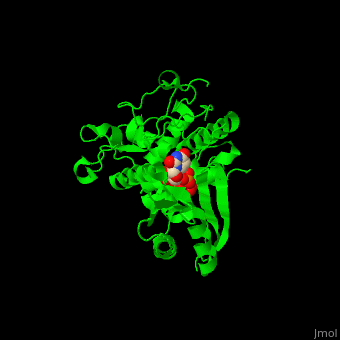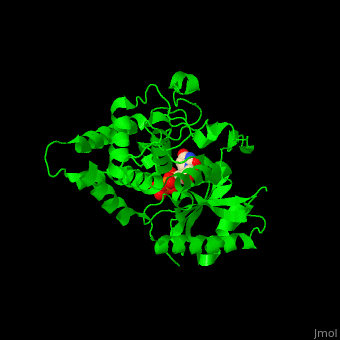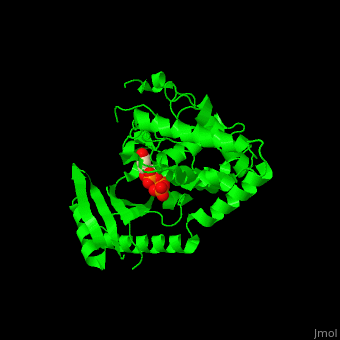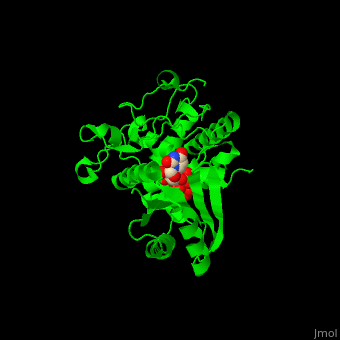
| - | '''Poly(A) RNA polymerase protein Cid1''' (Caffeine-Induced Death protein 1) is involved in cell cycle arrest where it inhibits unscheduled mitosis<ref>PMID:17072891</ref>. Cid1 is found in fission yeast. Cid1 exists in 3 forms: '''apo I''' consists of 4 subunits; '''apo II''' consists of 2 subunits; '''UTP-bound''' form consists of 4 subunits. | + | '''Poly(A) RNA polymerase protein Cid1''' (Caffeine-Induced Death protein 1) is involved in cell cycle arrest where it inhibits unscheduled mitosis<ref>PMID:17072891</ref>. Cid1 is found in fission yeast. Cid1 exists in 3 forms: '''apo I''' consists of 4 subunits; '''apo II''' consists of 2 subunits; '''UTP-bound''' form consists of 4 subunits. <br /> |
| + | '''Poly(A) RNA polymerase Gld2''' is responsible for poly(A) tail lengthening. Gld2 activity in the brain is required for long-term memory<ref>PMID:15987816</ref>. | ||
== Structural highlights == | == Structural highlights == | ||
Revision as of 08:04, 23 August 2018
| |||||||||||
3D Structures of poly(A) RNA polymerase protein Cid1
Updated on 23-August-2018
References
- ↑ Stevenson AL, Norbury CJ. The Cid1 family of non-canonical poly(A) polymerases. Yeast. 2006 Oct 15;23(13):991-1000. PMID:17072891 doi:http://dx.doi.org/10.1002/yea.1408
- ↑ Kachouri R, Stribinskis V, Zhu Y, Ramos KS, Westhof E, Li Y. A surprisingly large RNase P RNA in Candida glabrata. RNA. 2005 Jul;11(7):1064-72. doi: 10.1261/rna.2130705. PMID:15987816 doi:http://dx.doi.org/10.1261/rna.2130705
- ↑ Yates LA, Fleurdepine S, Rissland OS, De Colibus L, Harlos K, Norbury CJ, Gilbert RJ. Structural basis for the activity of a cytoplasmic RNA terminal uridylyl transferase. Nat Struct Mol Biol. 2012 Jul 1. doi: 10.1038/nsmb.2329. PMID:22751018 doi:10.1038/nsmb.2329
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Jaime Prilusky