This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Phage integrase
From Proteopedia
(Difference between revisions)
| (3 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
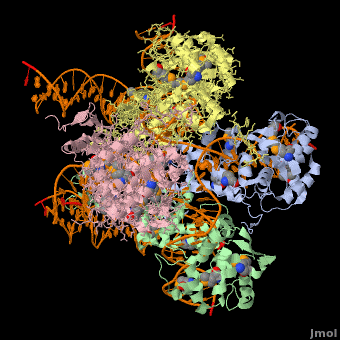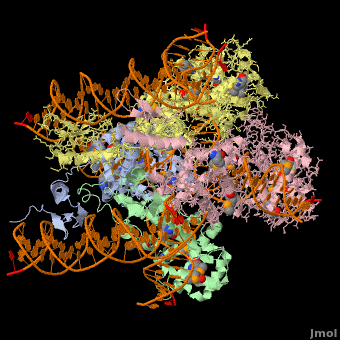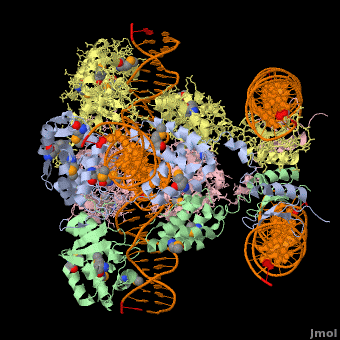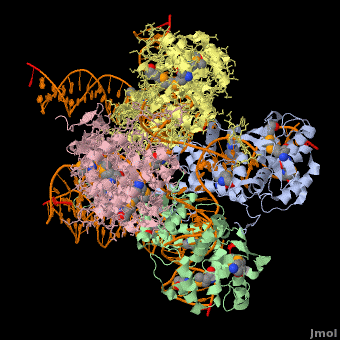
| - | <StructureSection load=' | + | <StructureSection load='' size='400' side='right' scene='44/443479/Cv/2' caption='Se-Met Enterobacteria phage λ integrase:DNA tetramer complex (Holiday Junction) [[1z1g]]' pspeed='8'> |
| - | + | ||
'''Phage integrase''' (PIN) mediates recombination between DNA recognition sequences - the phage attachment site attP and the bacterial attachment site attB<ref>PMID:14687564</ref>. The '''Tyrosine family PIN''' uses a catalytic tyrosine to mediate strand cleavage while the '''Serine family PIN''' uses a serine residue. The High-Pathogenicity Island (HPI) is a genomic island essential for virulence of the family of Enterobacteria. | '''Phage integrase''' (PIN) mediates recombination between DNA recognition sequences - the phage attachment site attP and the bacterial attachment site attB<ref>PMID:14687564</ref>. The '''Tyrosine family PIN''' uses a catalytic tyrosine to mediate strand cleavage while the '''Serine family PIN''' uses a serine residue. The High-Pathogenicity Island (HPI) is a genomic island essential for virulence of the family of Enterobacteria. | ||
| + | *The biological assembly of Enterobacteria phage λ integrase is <scene name='44/443479/Cv/3'>homotetramer</scene>. | ||
</StructureSection> | </StructureSection> | ||
| - | == 3D Structures of phage integrase == | + | == 3D Structures of phage integrase see [[Retroviral Integrase]]== |
| - | + | ||
| - | + | ||
| - | [[3nrw]] – PIN/site-specific recombinase N-terminal – ''Haloarcula marismortui''<br /> | ||
| - | [[3jtz]] – PIN arm-type binding domain – ''Yersinia pestis''<br /> | ||
| - | [[3ju0]] – HPI PIN arm-type binding domain – ''Pectobacterium atrosepticum''<br /> | ||
| - | [[2kkp]] – PIN SAM-like domain – ''Moorella thermoacetica'' – NMR<br /> | ||
| - | [[2kkv]] – PIN fragment – ''Salmonella enterica'' – NMR<br /> | ||
| - | [[2khq]] - PIN fragment – ''Staphylococcus saprophyticus'' – NMR<br /> | ||
| - | [[2wcc]] – λPIN DBD+ DNA – Enterobacteria phage λ - NMR<br /> | ||
| - | [[1kjk]] – λPIN N-terminal - NMR<br /> | ||
| - | [[2oxo]], [[1z19]], [[1z1b]] - λPIN core binding domain + DNA<br /> | ||
| - | [[1ae9]] - λPIN catalytic core domain (mutant) <br /> | ||
| - | [[1z1g]] – λPIN + DNA (Holliday junction)<br /> | ||
| - | [[1p7d]] – λPIN fragment + DNA <br /> | ||
| - | [[3bvp]] – PIN catalytic core domain – Lactococcus phage TP101-1<br /> | ||
| - | [[1aih]] - PIN catalytic core domain – Haemophilus phage HP1<br /> | ||
| - | [[2khv]], [[2kj5]] – PIN residues 102-199 – ''Nitrosospira multiformis'' - NMR | ||
== References == | == References == | ||
<references/> | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D Structures of phage integrase see Retroviral Integrase
References
- ↑ Groth AC, Calos MP. Phage integrases: biology and applications. J Mol Biol. 2004 Jan 16;335(3):667-78. PMID:14687564 doi:http://dx.doi.org/10.1016/S0022283603013561