This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
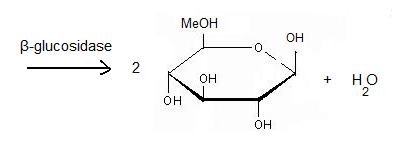
Beta-glucosidase
From Proteopedia
(Difference between revisions)
| Line 80: | Line 80: | ||
==Additional Resources== | ==Additional Resources== | ||
For additional information, see: [[Carbohydrate Metabolism]] | For additional information, see: [[Carbohydrate Metabolism]] | ||
| + | |||
| + | ==3D structures of Beta-glucosidase== | ||
| + | [[Beta-glucosidase 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
| Line 106: | Line 110: | ||
**[[1oif]], [[1od0]] – TmGB catalytic domain - ''Thermotoga maritima''<br /> | **[[1oif]], [[1od0]] – TmGB catalytic domain - ''Thermotoga maritima''<br /> | ||
**[[4gxp]] – TmGB/TrGB<br /> | **[[4gxp]] – TmGB/TrGB<br /> | ||
| - | **[[1ug6]] – | + | **[[1ug6]] – TtGB - ''Thermus thermophilus''<br /> |
| + | **[[4bce]] – TtGB (mutant) <br /> | ||
**[[1hxj]], [[1e1e]] – ZmGB – ''Zea mays''<br /> | **[[1hxj]], [[1e1e]] – ZmGB – ''Zea mays''<br /> | ||
**[[1e4l]] - ZmGB (mutant) <br /> | **[[1e4l]] - ZmGB (mutant) <br /> | ||
| Line 113: | Line 118: | ||
**[[1tr1]] - BpGBB (mutant) - ''Bacillus polymyxa''<br /> | **[[1tr1]] - BpGBB (mutant) - ''Bacillus polymyxa''<br /> | ||
**[[1cbg]] – GB cyanogenic – White clover<br /> | **[[1cbg]] – GB cyanogenic – White clover<br /> | ||
| - | **[[3aiu]] - | + | **[[3aiu]] - ryGB residues 50-568 – rye<br /> |
**[[3ut0]], [[3usz]] – PsGB residues 28-840 – ''Pseudoalteromonas''<br /> | **[[3ut0]], [[3usz]] – PsGB residues 28-840 – ''Pseudoalteromonas''<br /> | ||
**[[3f95]] – PsGB residues 657-840<br /> | **[[3f95]] – PsGB residues 657-840<br /> | ||
| Line 122: | Line 127: | ||
**[[3zyz]], [[3zz1]] – GB – ''Hypocrea jecorina''<br /> | **[[3zyz]], [[3zz1]] – GB – ''Hypocrea jecorina''<br /> | ||
**[[4hz6]] – ubGB – uncultured bacterium<br /> | **[[4hz6]] – ubGB – uncultured bacterium<br /> | ||
| - | **[[4bce]] – GB (mutant) – ''Thermus thermophilus''<br /> | ||
**[[4iib]] – AaGB – ''Aspergillus aculeatus''<br /> | **[[4iib]] – AaGB – ''Aspergillus aculeatus''<br /> | ||
**[[5fjj]] – GB – ''Aspergillus oryzae''<br /> | **[[5fjj]] – GB – ''Aspergillus oryzae''<br /> | ||
| Line 133: | Line 137: | ||
**[[3w53]] – GB – ''Micrococcus antarcticus''<br /> | **[[3w53]] – GB – ''Micrococcus antarcticus''<br /> | ||
**[[4mdo]] – HgGB – ''Humicola grisea''<br /> | **[[4mdo]] – HgGB – ''Humicola grisea''<br /> | ||
| - | **[[3wh5]], [[3u48]], [[3u4a]], [[5gnx]], [[5gny]], [[5gnz]], [[5wka]] – MeGB – ''Metagenome''<br /> | + | **[[3wh5]], [[3u48]], [[3u4a]], [[5gnx]], [[5gny]], [[5gnz]], [[5wka]], [[5ns6]] – MeGB – ''Metagenome''<br /> |
**[[5ayi]], [[5ayb]] – MeGB (mutant)<br /> | **[[5ayi]], [[5ayb]] – MeGB (mutant)<br /> | ||
**[[5ju6]] – GB – ''Talaromyces emersonii''<br /> | **[[5ju6]] – GB – ''Talaromyces emersonii''<br /> | ||
**[[5wug]], [[5wvp]] – GB – ''Paenibacillus barengoltzii''<br /> | **[[5wug]], [[5wvp]] – GB – ''Paenibacillus barengoltzii''<br /> | ||
**[[5xxl]] – BtGB – ''Bacterioides thetaiotaomicron''<br /> | **[[5xxl]] – BtGB – ''Bacterioides thetaiotaomicron''<br /> | ||
| + | **[[5ogz]] – GB – ''Ruminiclostridium thermocellum''<br /> | ||
*Beta-glucosidase complex with sugar | *Beta-glucosidase complex with sugar | ||
| Line 154: | Line 159: | ||
**[[3aht]], [[3ahv]], [[3scq]], [[3scs]], [[4qlk]], [[4qlj]] – JrGB7 (mutant) + saccharide<br /> | **[[3aht]], [[3ahv]], [[3scq]], [[3scs]], [[4qlk]], [[4qlj]] – JrGB7 (mutant) + saccharide<br /> | ||
**[[1oin]] - TmGBA + glucoside<br /> | **[[1oin]] - TmGBA + glucoside<br /> | ||
| + | **[[5oss]] – TmGB + glucoimidazole<br /> | ||
**[[4hz7]], [[4hz8]] - UbGB residues 18-482 (mutant) + glucoside<br /> | **[[4hz7]], [[4hz8]] - UbGB residues 18-482 (mutant) + glucoside<br /> | ||
**[[2zox]], [[2e9l]], [[2e9m]] - hGB cystolic + glucoside<br /> | **[[2zox]], [[2e9l]], [[2e9m]] - hGB cystolic + glucoside<br /> | ||
| Line 199: | Line 205: | ||
**[[1y7v]] - hGB (mutant) + inhibitor<br /> | **[[1y7v]] - hGB (mutant) + inhibitor<br /> | ||
**[[5bx3]] - TxGB + inhibitor<br /> | **[[5bx3]] - TxGB + inhibitor<br /> | ||
| + | **[[5ns8]] – MeGB (mutant) + deoxynojirimycin<br /> | ||
*6-phospho-β-glucosidase | *6-phospho-β-glucosidase | ||
| Line 205: | Line 212: | ||
**[[1up4]] – TmPGB <br /> | **[[1up4]] – TmPGB <br /> | ||
**[[1up6]], [[1up7]] – TmPGB + NAD + G6P<br /> | **[[1up6]], [[1up7]] – TmPGB + NAD + G6P<br /> | ||
| - | **[[3qom]], [[4gze]] – PGB – ''Lactobacillus plantarum''<br /> | + | **[[3qom]], [[4gze]], [[5nav]] – PGB – ''Lactobacillus plantarum''<br /> |
**[[2xhy]] – EcPGB <br /> | **[[2xhy]] – EcPGB <br /> | ||
**[[1h4p]] – PGB I/II – yeast<br /> | **[[1h4p]] – PGB I/II – yeast<br /> | ||
Revision as of 08:18, 8 April 2019
| |||||||||||
3D structures of Beta-glucosidase
Updated on 08-April-2019
References
- ↑ Aguilar M, Gloster TM, Garcia-Moreno MI, Ortiz Mellet C, Davies GJ, Llebaria A, Casas J, Egido-Gabas M, Garcia Fernandez JM. Molecular basis for beta-glucosidase inhibition by ring-modified calystegine analogues. Chembiochem. 2008 Nov 3;9(16):2612-8. PMID:18833549 doi:10.1002/cbic.200800451
- ↑ http://en.wikipedia.org/wiki/B-glucosidase
- ↑ Davies G, Henrissat B. Structures and mechanisms of glycosyl hydrolases. Structure. 1995 Sep 15;3(9):853-9. PMID:8535779
- ↑ http://www.ebi.ac.uk/interpro/IEntry?ac=IPR018120#PUB00002205
- ↑ http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/CSA/CSA_Site_Wrapper.pl?pdb=2vrj
- ↑ Davies G, Henrissat B. Structures and mechanisms of glycosyl hydrolases. Structure. 1995 Sep 15;3(9):853-9. PMID:8535779
- ↑ http://www.cazy.org/fam/ghf_INV_RET.html#3
- ↑ Dvir H, Harel M, McCarthy AA, Toker L, Silman I, Futerman AH, Sussman JL. X-ray structure of human acid-beta-glucosidase, the defective enzyme in Gaucher disease. EMBO Rep. 2003 Jul;4(7):704-9. PMID:12792654 doi:10.1038/sj.embor.embor873
- ↑ 9.0 9.1 9.2 Premkumar L, Sawkar AR, Boldin-Adamsky S, Toker L, Silman I, Kelly JW, Futerman AH, Sussman JL. X-ray structure of human acid-beta-glucosidase covalently bound to conduritol-B-epoxide. Implications for Gaucher disease. J Biol Chem. 2005 Jun 24;280(25):23815-9. Epub 2005 Apr 6. PMID:15817452 doi:M502799200
- ↑ Hrmova M, Varghese JN, De Gori R, Smith BJ, Driguez H, Fincher GB. Catalytic mechanisms and reaction intermediates along the hydrolytic pathway of a plant beta-D-glucan glucohydrolase. Structure. 2001 Nov;9(11):1005-16. PMID:11709165
- ↑ Zeev-Ben-Mordehai T, Silman I, Sussman JL. Acetylcholinesterase in motion: visualizing conformational changes in crystal structures by a morphing procedure. Biopolymers. 2003 Mar;68(3):395-406. PMID:12601798 doi:10.1002/bip.10287
- ↑ 12.0 12.1 12.2 12.3 Shaaltiel Y, Bartfeld D, Hashmueli S, Baum G, Brill-Almon E, Galili G, Dym O, Boldin-Adamsky SA, Silman I, Sussman JL, Futerman AH, Aviezer D. Production of glucocerebrosidase with terminal mannose glycans for enzyme replacement therapy of Gaucher's disease using a plant cell system. Plant Biotechnol J. 2007 Sep;5(5):579-90. Epub 2007 May 24. PMID:17524049 doi:10.1111/j.1467-7652.2007.00263.x
- ↑ Brumshtein B, Greenblatt HM, Butters TD, Shaaltiel Y, Aviezer D, Silman I, Futerman AH, Sussman JL. Crystal structures of complexes of N-butyl- and N-nonyl-deoxynojirimycin bound to acid beta-glucosidase: insights into the mechanism of chemical chaperone action in Gaucher disease. J Biol Chem. 2007 Sep 28;282(39):29052-8. Epub 2007 Jul 31. PMID:17666401 doi:10.1074/jbc.M705005200
- ↑ Lieberman RL, Wustman BA, Huertas P, Powe AC Jr, Pine CW, Khanna R, Schlossmacher MG, Ringe D, Petsko GA. Structure of acid beta-glucosidase with pharmacological chaperone provides insight into Gaucher disease. Nat Chem Biol. 2007 Feb;3(2):101-7. Epub 2006 Dec 24. PMID:17187079 doi:http://dx.doi.org/10.1038/nchembio850
- ↑ Brumshtein B, Wormald MR, Silman I, Futerman AH, Sussman JL. Structural comparison of differently glycosylated forms of acid-beta-glucosidase, the defective enzyme in Gaucher disease. Acta Crystallogr D Biol Crystallogr. 2006 Dec;62(Pt 12):1458-65. Epub 2006, Nov 23. PMID:17139081 doi:S0907444906038303
- ↑ 16.0 16.1 Brumshtein B, Salinas P, Peterson B, Chan V, Silman I, Sussman JL, Savickas PJ, Robinson GS, Futerman AH. Characterization of gene-activated human acid-beta-glucosidase: crystal structure, glycan composition, and internalization into macrophages. Glycobiology. 2010 Jan;20(1):24-32. Epub 2009 Sep 9. PMID:19741058 doi:10.1093/glycob/cwp138
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Muriel Breteau, Alexander Berchansky, Joel L. Sussman, David Canner