This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hepatocyte growth factor activator
From Proteopedia
(Difference between revisions)
| (7 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
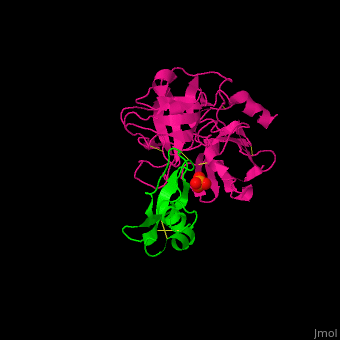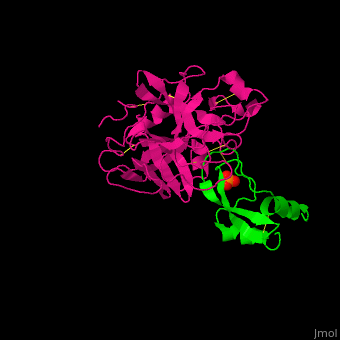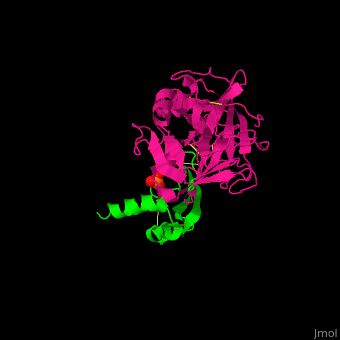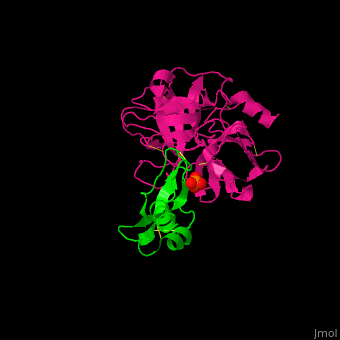
| - | <StructureSection load=' | + | <StructureSection load='' size='350' side='right' caption='Human hepatocyte growth factor activator (magenta) complex with HGFA inhibitor type 1 (green) and phosphate (PDB code [[1yc0]])' scene='70/706236/Cv/1'> |
== Function == | == Function == | ||
'''Hepatocyte growth factor activator''' (HGFA) is a serine protease which links tissue injury to activation of hepatocyte growth factor (HGF). HGFA converts HGF from its latent single-chain form to a heterodimeric active form. HGFA is synthesized as an inactive precursor which is converted to a heterodimer via proteolysis as a result of tissue injury<ref>PMID:20402766</ref>. | '''Hepatocyte growth factor activator''' (HGFA) is a serine protease which links tissue injury to activation of hepatocyte growth factor (HGF). HGFA converts HGF from its latent single-chain form to a heterodimeric active form. HGFA is synthesized as an inactive precursor which is converted to a heterodimer via proteolysis as a result of tissue injury<ref>PMID:20402766</ref>. | ||
| - | |||
| - | == Disease == | ||
== Relevance == | == Relevance == | ||
| Line 9: | Line 7: | ||
== Structural highlights == | == Structural highlights == | ||
| - | + | HGFA substrate binding cleft resembles other trypsin-like serine proteases with <scene name='70/706236/Cv/4'>inhibitors of the Kunitz type (α/β folds rich in Cys-Cys bonds)</scene> <ref>PMID:157134585</ref>. | |
</StructureSection> | </StructureSection> | ||
| Line 22: | Line 20: | ||
== References == | == References == | ||
<references/> | <references/> | ||
| + | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D Structures of hepatocyte growth factor activator
Updated on 15-May-2019
1ybw – hHGFA protease domain – human
1yc0 – hHGFA protease domain + Kunitz-type protease inhibitor
2r0l, 2r0k, 2wub, 2wuc, 3k2u – hHGFA protease domain + antibody
References
- ↑ Miyazawa K. Hepatocyte growth factor activator (HGFA): a serine protease that links tissue injury to activation of hepatocyte growth factor. FEBS J. 2010 May;277(10):2208-14. doi: 10.1111/j.1742-4658.2010.07637.x. Epub, 2010 Apr 2. PMID:20402766 doi:http://dx.doi.org/10.1111/j.1742-4658.2010.07637.x
- ↑ Wu KL, Li YQ, Tabassum A, Lu WY, Aubert I, Wong CS. Loss of neuronal protein expression in mouse hippocampus after irradiation. J Neuropathol Exp Neurol. 2010 Mar;69(3):272-80. doi:, 10.1097/NEN.0b013e3181d1afe4. PMID:20142763 doi:http://dx.doi.org/10.1097/NEN.0b013e3181d1afe4
- ↑ . PMID:157134585