This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Exonuclease
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
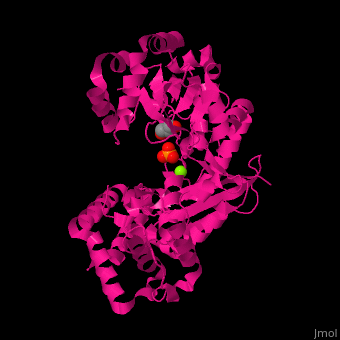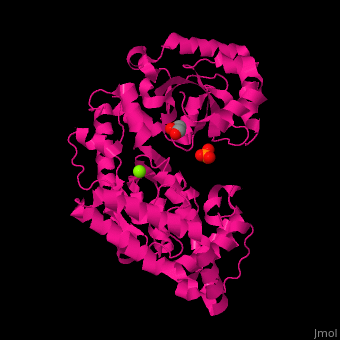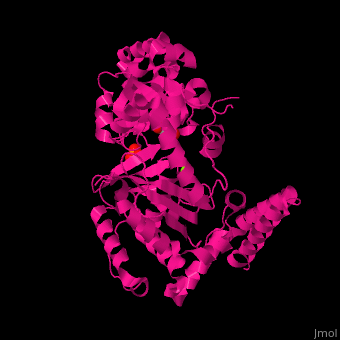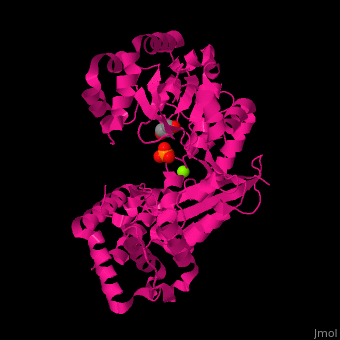
<scene name='46/466467/Cv/3'>Mg coordination site</scene> in ''E. coli'' exonuclease I (PDB code [[1fxx]]).<ref>PMID:11101894</ref> Water molecules shown as red spheres. | <scene name='46/466467/Cv/3'>Mg coordination site</scene> in ''E. coli'' exonuclease I (PDB code [[1fxx]]).<ref>PMID:11101894</ref> Water molecules shown as red spheres. | ||
| + | |||
| + | == 3D Structures of exonuclease == | ||
| + | [[Exonuclease 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
| Line 22: | Line 26: | ||
*ExN-I | *ExN-I | ||
| + | **[[3qea]], [[3qeb]], [[5v04]], [[5v05]], [[5v06]], [[5v0b]], [[5v0c]], [[5v0d]], [[5v0e]], [[5uzv]] - hExN-I + DNA - human<br /> | ||
| + | **[[3qe9]], [[5v07]], [[5v08]], [[5v09]], [[5v0a]] – hExN-I (mutant) + DNA – human<br /> | ||
**[[1fxx]], [[3c95]] – EcExN-I – ''Escherichia coli''<br /> | **[[1fxx]], [[3c95]] – EcExN-I – ''Escherichia coli''<br /> | ||
**[[2qxf]] - EcExN-I + TMP<br /> | **[[2qxf]] - EcExN-I + TMP<br /> | ||
| Line 27: | Line 33: | ||
**[[3c94]] - EcExN-I + DNA-binding C terminal peptide<br /> | **[[3c94]] - EcExN-I + DNA-binding C terminal peptide<br /> | ||
**[[4jrp]], [[4jrq]], [[4js5]], [[4js4]], [[4hcc]], [[4hcb]] – EcExN-I + DNA<br /> | **[[4jrp]], [[4jrq]], [[4js5]], [[4js4]], [[4hcc]], [[4hcb]] – EcExN-I + DNA<br /> | ||
| - | **[[3qe9]], [[5v07]], [[5v08]], [[5v09]], [[5v0a]] – hExN-I (mutant) + DNA – human<br /> | ||
| - | **[[3qea]], [[3qeb]], [[5v04]], [[5v05]], [[5v06]], [[5v0b]], [[5v0c]], [[5v0d]], [[5v0e]], [[5uzv] - hExN-I + DNA<br /> | ||
**[[4rg8]] – ExN-I (mutant) – ''Methylocaldum szegedienes''<br /> | **[[4rg8]] – ExN-I (mutant) – ''Methylocaldum szegedienes''<br /> | ||
**[[5fiq]], [[5fis]] – ExN-I ExN domain – silkworm<br /> | **[[5fiq]], [[5fis]] – ExN-I ExN domain – silkworm<br /> | ||
| Line 89: | Line 93: | ||
*3'-5' ExN | *3'-5' ExN | ||
| + | **[[4qoz]], [[4l8r]] – hExN + histone RNA hairpin-binding protein + RNA<br /> | ||
**[[4yor]], [[4yot]], [[4you]] – PhExN phoexo I – ''Pyrococcus horikoshii''<br /> | **[[4yor]], [[4yot]], [[4you]] – PhExN phoexo I – ''Pyrococcus horikoshii''<br /> | ||
**[[4yov]], [[4yow]], [[4yox]], [[4yoy]] – PhExN phoexo I (mutant) + DNA<br /> | **[[4yov]], [[4yow]], [[4yox]], [[4yoy]] – PhExN phoexo I (mutant) + DNA<br /> | ||
| - | **[[4qoz]], [[4l8r]] – hExN + histone RNA hairpin-binding protein + RNA<br /> | ||
*5’-3’ ExN | *5’-3’ ExN | ||
| Line 104: | Line 108: | ||
**[[3mxj]] - mExN Trex1<br /> | **[[3mxj]] - mExN Trex1<br /> | ||
**[[2o4i]], [[3mxi]], [[3mxm]], [[3u3y]], [[3u6f]] - mExN Trex1 (mutant) + polynucleotide<br /> | **[[2o4i]], [[3mxi]], [[3mxm]], [[3u3y]], [[3u6f]] - mExN Trex1 (mutant) + polynucleotide<br /> | ||
| - | **[[2oa8]] - mExN Trex1 + DNA<br /> | + | **[[2oa8]], [[5ywv]], [[5ywu]], [[5ywt]], [[5yws]] - mExN Trex1 + DNA<br /> |
**[[4ynq]] - mExN Trex1 (mutant) + DNA<br /> | **[[4ynq]] - mExN Trex1 (mutant) + DNA<br /> | ||
**[[3b6o]], [[3b6p]] - mExN Trex1 + ion inhibitor<br /> | **[[3b6o]], [[3b6p]] - mExN Trex1 + ion inhibitor<br /> | ||
| + | **[[6a45]] - mExN Trex2<br /> | ||
| + | **[[6a4b]], [[6a47]] - mExN Trex2 + DNA<br /> | ||
| + | **[[6a46]] - mExN Trex2 + nucleotide<br /> | ||
*Other ExN | *Other ExN | ||
| - | **[[1ir6]], [[2zxo]], [[2zxp]], [[2zxr]] – ExN Recj – ''Thermus thermophiles''<br /> | ||
**[[1w0h]] – hExn Eri1 nuclease domain <br /> | **[[1w0h]] – hExn Eri1 nuclease domain <br /> | ||
**[[1zbu]] - hExn Eri1<br /> | **[[1zbu]] - hExn Eri1<br /> | ||
**[[1zbh]] - hExn Eri1 (mutant) + DNA<br /> | **[[1zbh]] - hExn Eri1 (mutant) + DNA<br /> | ||
**[[5aho]] – hExN Apollo<br /> | **[[5aho]] – hExN Apollo<br /> | ||
| + | **[[5zyw]] – hExN mitochondrial genome repair 1<br /> | ||
| + | **[[5zyv]], [[5zyu]], [[5zyt]] – hExN mitochondrial genome repair 1 + DNA<br /> | ||
| + | **[[3e2v]] – yExN <br /> | ||
**[[2w45]] – H4ExN alkaline - Herpesvirus 4<br /> | **[[2w45]] – H4ExN alkaline - Herpesvirus 4<br /> | ||
**[[2w4b]] - H4ExN alkaline (mutant)<br /> | **[[2w4b]] - H4ExN alkaline (mutant)<br /> | ||
| Line 120: | Line 129: | ||
**[[3sz4]] - LhExN alkaline + AMP<br /> | **[[3sz4]] - LhExN alkaline + AMP<br /> | ||
**[[3sz5]] - LhExN alkaline + polynucleotide<br /> | **[[3sz5]] - LhExN alkaline + polynucleotide<br /> | ||
| - | **[[3e2v]] – yExN <br /> | ||
**[[4lty]], [[4lu9]], [[4m0v]] – EcExN sbcD<br /> | **[[4lty]], [[4lu9]], [[4m0v]] – EcExN sbcD<br /> | ||
| + | **[[6m9k]] - ExN + recombination protein BET – Escherichia phage λ<br /> | ||
| + | **[[1ir6]], [[2zxo]], [[2zxp]], [[2zxr]] – ExN Recj – ''Thermus thermophiles''<br /> | ||
}} | }} | ||
Revision as of 07:54, 25 June 2019
| |||||||||||
3D Structures of exonuclease
Updated on 25-June-2019
References
- ↑ Mukherjee D, Fritz DT, Kilpatrick WJ, Gao M, Wilusz J. Analysis of RNA exonucleolytic activities in cellular extracts. Methods Mol Biol. 2004;257:193-212. PMID:14770007 doi:http://dx.doi.org/10.1385/1-59259-750-5:193
- ↑ Breyer WA, Matthews BW. Structure of Escherichia coli exonuclease I suggests how processivity is achieved. Nat Struct Biol. 2000 Dec;7(12):1125-8. PMID:11101894 doi:10.1038/81978