This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
S100 protein
From Proteopedia
(Difference between revisions)
| Line 29: | Line 29: | ||
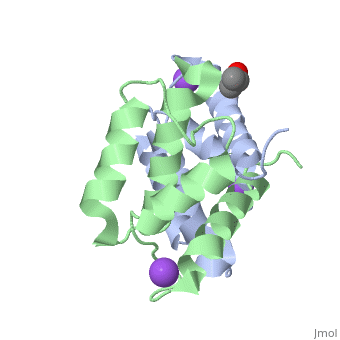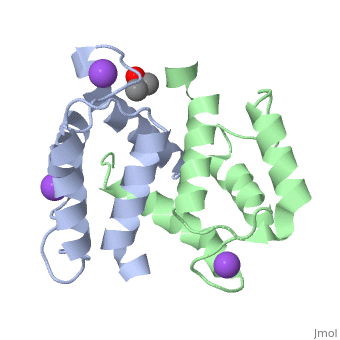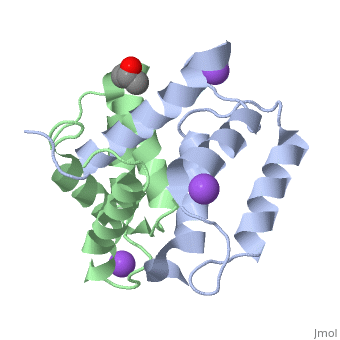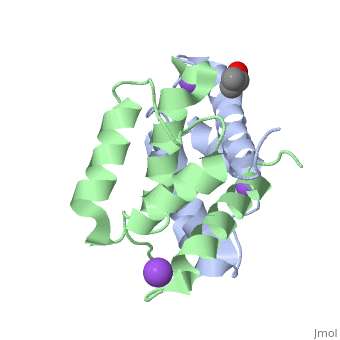
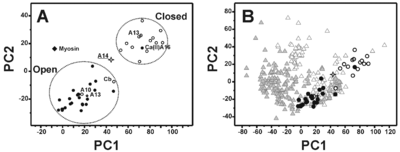
S100A16 is a special member of the S100 class of calcium binding proteins, because it <scene name='Journal:JBIC:3/As/12'>performs a conformational change upon calcium(II) binding</scene> much smaller than experienced by most S100 proteins. This was observed after determination of the solution structures of apo and <scene name='Journal:JBIC:3/Dual_binding_calcium/3'>calcium(II)-bound S100A16</scene> and the <scene name='Journal:JBIC:3/Crysal/2'>crystal structure of apo S100A16</scene>. The likely reason for minimal conformational change <scene name='Journal:JBIC:3/Calcium_binding_start/7'>in S100A16</scene> is the lower calcium binding affinity and stronger <scene name='Journal:JBIC:3/Hydrophobic_interactions_2/3'>hydrophobic interaction</scene> between <scene name='Journal:JBIC:3/Please_work/3'>helix III and IV present in this protein </scene> with respect to other S100 proteins. Another characteristic of <scene name='Journal:JBIC:3/Opening/3'>S100A16</scene> is that the helix IV has the same length in <scene name='Journal:JBIC:3/25_residue_long_apo/3'>both apo</scene> and <scene name='Journal:JBIC:3/25_residue_calclium_bound/3'>calcium(II) forms</scene> because of <scene name='Journal:JBIC:3/Motif_good/5'>the presence of a Gly-Gly-Ile-Thr-Gly-Pro sequence motif</scene> in helix IV. Based on the available structures of S100 members, we analyzed and summarized all their conformational changes due to calcium(II) binding by a principal component analysis. <scene name='Journal:JBIC:3/Calcium_binding_start/7'>Calcium binding</scene> was proved by both NMR titration and Isothermal Titration Calorimetry (ITC) experiments. Even if the <scene name='Journal:JBIC:3/Binding_calcium_glu/2'>important Glu residue</scene> in the last position of first EF-hand calcium binding loop <scene name='Journal:JBIC:3/Binding_calcium/13'>is missing</scene>, these experimental data indicated that S100A16 can still bind one calcium(II) ion in such loop. NMR relaxation <scene name='Journal:JBIC:3/Flexible_broadwide/4'>studies showed that the first calcium binding loop and the beginning of the second helix</scene> are the most <scene name='Journal:JBIC:3/Flexible_broad/3'>flexible regions in both the apo and calcium(II)-bound S100A16</scene>. Although the biological function of S100A16 is still unclear yet, these structural and dynamic properties can provide useful information for further functional studies. | S100A16 is a special member of the S100 class of calcium binding proteins, because it <scene name='Journal:JBIC:3/As/12'>performs a conformational change upon calcium(II) binding</scene> much smaller than experienced by most S100 proteins. This was observed after determination of the solution structures of apo and <scene name='Journal:JBIC:3/Dual_binding_calcium/3'>calcium(II)-bound S100A16</scene> and the <scene name='Journal:JBIC:3/Crysal/2'>crystal structure of apo S100A16</scene>. The likely reason for minimal conformational change <scene name='Journal:JBIC:3/Calcium_binding_start/7'>in S100A16</scene> is the lower calcium binding affinity and stronger <scene name='Journal:JBIC:3/Hydrophobic_interactions_2/3'>hydrophobic interaction</scene> between <scene name='Journal:JBIC:3/Please_work/3'>helix III and IV present in this protein </scene> with respect to other S100 proteins. Another characteristic of <scene name='Journal:JBIC:3/Opening/3'>S100A16</scene> is that the helix IV has the same length in <scene name='Journal:JBIC:3/25_residue_long_apo/3'>both apo</scene> and <scene name='Journal:JBIC:3/25_residue_calclium_bound/3'>calcium(II) forms</scene> because of <scene name='Journal:JBIC:3/Motif_good/5'>the presence of a Gly-Gly-Ile-Thr-Gly-Pro sequence motif</scene> in helix IV. Based on the available structures of S100 members, we analyzed and summarized all their conformational changes due to calcium(II) binding by a principal component analysis. <scene name='Journal:JBIC:3/Calcium_binding_start/7'>Calcium binding</scene> was proved by both NMR titration and Isothermal Titration Calorimetry (ITC) experiments. Even if the <scene name='Journal:JBIC:3/Binding_calcium_glu/2'>important Glu residue</scene> in the last position of first EF-hand calcium binding loop <scene name='Journal:JBIC:3/Binding_calcium/13'>is missing</scene>, these experimental data indicated that S100A16 can still bind one calcium(II) ion in such loop. NMR relaxation <scene name='Journal:JBIC:3/Flexible_broadwide/4'>studies showed that the first calcium binding loop and the beginning of the second helix</scene> are the most <scene name='Journal:JBIC:3/Flexible_broad/3'>flexible regions in both the apo and calcium(II)-bound S100A16</scene>. Although the biological function of S100A16 is still unclear yet, these structural and dynamic properties can provide useful information for further functional studies. | ||
| + | |||
| + | == 3D Structures of S100 proteins == | ||
| + | [[S100 proteins 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
== 3D Structures of S100 proteins == | == 3D Structures of S100 proteins == | ||
| Line 37: | Line 41: | ||
*S100-A1 | *S100-A1 | ||
| - | **[[1k2h]] – rCBP-1 – rat - NMR <br /> | ||
| - | **[[2k2f]] – rCBP-1 + Ca + ryanodine receptor peptide - NMR<br /> | ||
| - | **[[2kbm]] – rCBP-1 + F-actin capping protein peptide - NMR<br /> | ||
**[[1zfs]], [[2lp2]], [[2lp3]], [[2l0p]] – hCBP-1 + Ca – human - NMR<br /> | **[[1zfs]], [[2lp2]], [[2lp3]], [[2l0p]] – hCBP-1 + Ca – human - NMR<br /> | ||
**[[5k89]] – hCBP-1 + Ca <br /> | **[[5k89]] – hCBP-1 + Ca <br /> | ||
| Line 46: | Line 47: | ||
**[[2llt]], [[2llu]] – hCBP-1 - NMR<br /> | **[[2llt]], [[2llu]] – hCBP-1 - NMR<br /> | ||
**[[2jpt]] – bCBP-1 (mutant) – bovine - NMR <br /> | **[[2jpt]] – bCBP-1 (mutant) – bovine - NMR <br /> | ||
| + | **[[1k2h]] – rCBP-1 – rat - NMR <br /> | ||
| + | **[[2k2f]] – rCBP-1 + Ca + ryanodine receptor peptide - NMR<br /> | ||
| + | **[[2kbm]] – rCBP-1 + F-actin capping protein peptide - NMR<br /> | ||
*S100-A2 | *S100-A2 | ||
| Line 76: | Line 80: | ||
*S100-A6 (Calcyclin) | *S100-A6 (Calcyclin) | ||
| - | **[[1cnp]], [[1a03]], [[2cnp]], [[1jwd]] – raCBP-6 – rabbit - NMR<br /> | ||
| - | **[[2jtt]] – raCBP-6 + calcyclin-binding protein - NMR<br /> | ||
**[[1kso]], [[1k8u]], [[1k9p]] – hCBP-6 <br /> | **[[1kso]], [[1k8u]], [[1k9p]] – hCBP-6 <br /> | ||
**[[1k96]], [[1k9k]] – hCBP-6 + Ca <br /> | **[[1k96]], [[1k9k]] – hCBP-6 + Ca <br /> | ||
**[[2m1k]] – hCBP-6 (mutant) + RAGE receptor - NMR <br /> | **[[2m1k]] – hCBP-6 (mutant) + RAGE receptor - NMR <br /> | ||
**[[4p2y]], [[4ybh]] – hCBP-6 + RAGE receptor + Ca + Zn <br /> | **[[4p2y]], [[4ybh]] – hCBP-6 + RAGE receptor + Ca + Zn <br /> | ||
| + | **[[1cnp]], [[1a03]], [[2cnp]], [[1jwd]] – raCBP-6 – rabbit - NMR<br /> | ||
| + | **[[2jtt]] – raCBP-6 + calcyclin-binding protein - NMR<br /> | ||
*S100-A7 (Psoriasin) | *S100-A7 (Psoriasin) | ||
| Line 98: | Line 102: | ||
**[[5w1f]] – hCBP-8 (mutant) + hCBP-9 (mutant) + Ni + Ca <br /> | **[[5w1f]] – hCBP-8 (mutant) + hCBP-9 (mutant) + Ni + Ca <br /> | ||
**[[4xjk]] – hCBP-8 (mutant) + hCBP-9 (mutant) + Ca + Mn <br /> | **[[4xjk]] – hCBP-8 (mutant) + hCBP-9 (mutant) + Ca + Mn <br /> | ||
| - | |||
*S100-A9 (Calgranulin-B or Migration inhibitory factor-related protein 14) | *S100-A9 (Calgranulin-B or Migration inhibitory factor-related protein 14) | ||
| Line 122: | Line 125: | ||
*S100-A11 (Calgizzarin) | *S100-A11 (Calgizzarin) | ||
| - | **[[1qls]] – CBP-11 + Ca + annexin I - pig <br /> | ||
| - | **[[1nsh]] – raCBP-11 - NMR<br /> | ||
**[[2luc]] – hCBP-11 - NMR<br /> | **[[2luc]] – hCBP-11 - NMR<br /> | ||
**[[1v4z]], [[1v50]] – hCBP-11 N terminal - NMR<br /> | **[[1v4z]], [[1v50]] – hCBP-11 N terminal - NMR<br /> | ||
| + | **[[1qls]] – pCBP-11 + Ca + annexin I - pig <br /> | ||
| + | **[[1nsh]] – raCBP-11 - NMR<br /> | ||
*S100-A12 | *S100-A12 | ||
| Line 138: | Line 141: | ||
*S100-A13 | *S100-A13 | ||
| - | **[[2cxj]] – CBP-13 – mouse - NMR<br /> | ||
**[[1yur]], [[1yus]] – hCBP-13 - NMR<br /> | **[[1yur]], [[1yus]] – hCBP-13 - NMR<br /> | ||
**[[1yut]], [[1yuu]] – hCBP-13 + Ca - NMR<br /> | **[[1yut]], [[1yuu]] – hCBP-13 + Ca - NMR<br /> | ||
| Line 148: | Line 150: | ||
**[[2le9]] – hCBP-13 + RAGEC2 - NMR<br /> | **[[2le9]] – hCBP-13 + RAGEC2 - NMR<br /> | ||
**[[2kot]] – hCBP-13 + amlexanox - NMR<br /> | **[[2kot]] – hCBP-13 + amlexanox - NMR<br /> | ||
| + | **[[2cxj]] – CBP-13 – mouse - NMR<br /> | ||
*S100-A14 | *S100-A14 | ||
| Line 165: | Line 168: | ||
*S100B | *S100B | ||
| - | **[[1sym]], [[1b4c]] – rCBP – rat - NMR<br /> | ||
| - | **[[1qlk]], [[2k7o]] – rCBP + Ca - NMR<br /> | ||
| - | **[[1xyd]] – rCBP + Zn + Ca - NMR<br /> | ||
| - | **[[1dt7]] – rCBP + Ca + p53 peptide - NMR<br /> | ||
| - | **[[1mwn]] – rCBP + F-actin capping protein peptide - NMR<br /> | ||
**[[1mq1]] – hCBP + F-actin capping protein peptide - NMR<br /> | **[[1mq1]] – hCBP + F-actin capping protein peptide - NMR<br /> | ||
| + | **[[1uwo]], [[2pru]] – hCBP - NMR <br /> | ||
| + | **[[2h61]], [[3hcm]] – hCBP + Ca <br /> | ||
| + | **[[4xyn]], [[5d7f]] – hCBP + Ca + RAGE peptide <br /> | ||
| + | **[[3czt]], [[3d0y]], [[3d10]] – hCBP + Zn + Ca <br /> | ||
| + | **[[2m49]] – hCBP + fibroblast growth factor 2 - NMR<br /> | ||
| + | **[[5csi]], [[5csj]], [[5csn]], [[5csf]] – hCBP subunit β + ribosomal protein S6 kinase α-1 peptide + Ca <br /> | ||
**[[3rm1]] – bCBP + F-actin capping protein peptide + Ca<br /> | **[[3rm1]] – bCBP + F-actin capping protein peptide + Ca<br /> | ||
**[[1psb]] – bCBP + NDR kinase peptide - NMR<br /> | **[[1psb]] – bCBP + NDR kinase peptide - NMR<br /> | ||
| Line 179: | Line 183: | ||
**[[4pdz]], [[4pe0]], [[4pe1]], [[4pe4]], [[4pe7]], [[5dkn]], [[5dkq]], [[5dkr]], [[5er4]], [[5er5]] – bCBP + Ca + inhibitor<br /> | **[[4pdz]], [[4pe0]], [[4pe1]], [[4pe4]], [[4pe7]], [[5dkn]], [[5dkq]], [[5dkr]], [[5er4]], [[5er5]] – bCBP + Ca + inhibitor<br /> | ||
**[[4fqo]] – bCBP (mutant) + Ca + inhibitor<br /> | **[[4fqo]] – bCBP (mutant) + Ca + inhibitor<br /> | ||
| - | **[[ | + | **[[1sym]], [[1b4c]] – rCBP – rat - NMR<br /> |
| - | **[[ | + | **[[1qlk]], [[2k7o]] – rCBP + Ca - NMR<br /> |
| - | + | **[[1xyd]] – rCBP + Zn + Ca - NMR<br /> | |
| - | **[[ | + | **[[1dt7]] – rCBP + Ca + p53 peptide - NMR<br /> |
| - | **[[ | + | **[[1mwn]] – rCBP + F-actin capping protein peptide - NMR<br /> |
| - | **[[ | + | |
*Calbindin D9k (S100G) | *Calbindin D9k (S100G) | ||
Revision as of 11:48, 31 December 2019
| |||||||||||
3D Structures of S100 proteins
Updated on 31-December-2019
References
- ↑ Donato R, Cannon BR, Sorci G, Riuzzi F, Hsu K, Weber DJ, Geczy CL. Functions of S100 proteins. Curr Mol Med. 2013 Jan;13(1):24-57. PMID:22834835
- ↑ Yao R, Lopez-Beltran A, Maclennan GT, Montironi R, Eble JN, Cheng L. Expression of S100 protein family members in the pathogenesis of bladder tumors. Anticancer Res. 2007 Sep-Oct;27(5A):3051-8. PMID:17970044
- ↑ Wilsmann-Theis D, Wagenpfeil J, Holzinger D, Roth J, Koch S, Schnautz S, Bieber T, Wenzel J. Among the S100 proteins, S100A12 is the most significant marker for psoriasis disease activity. J Eur Acad Dermatol Venereol. 2016 Jul;30(7):1165-70. doi: 10.1111/jdv.13269., Epub 2015 Sep 2. PMID:26333514 doi:http://dx.doi.org/10.1111/jdv.13269
- ↑ Zhu L, Okano S, Takahara M, Chiba T, Tu Y, Oda Y, Furue M. Expression of S100 protein family members in normal skin and sweat gland tumors. J Dermatol Sci. 2013 Jun;70(3):211-9. doi: 10.1016/j.jdermsci.2013.03.002. Epub, 2013 Mar 16. PMID:23623205 doi:http://dx.doi.org/10.1016/j.jdermsci.2013.03.002
- ↑ Bertini I, Borsi V, Cerofolini L, Das Gupta S, Fragai M, Luchinat C. Solution structure and dynamics of human S100A14. J Biol Inorg Chem. 2012 Nov 30. PMID:23197251 doi:10.1007/s00775-012-0963-3
- ↑ Babini E, Bertini I, Borsi V, Calderone V, Hu X, Luchinat C, Parigi G. Structural characterization of human S100A16, a low-affinity calcium binder. J Biol Inorg Chem. 2010 Nov 3. PMID:21046186 doi:10.1007/s00775-010-0721-3