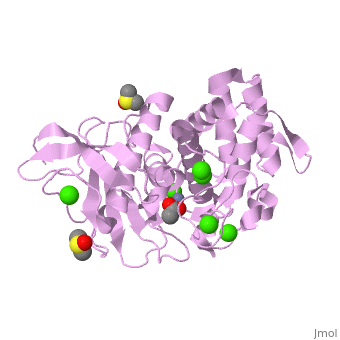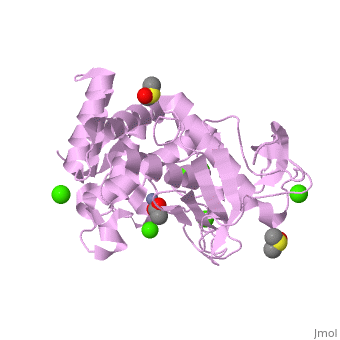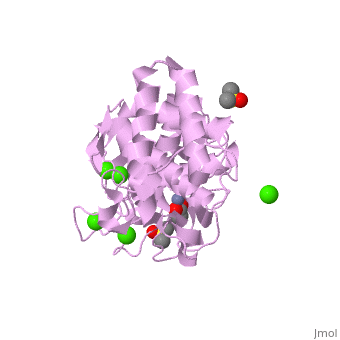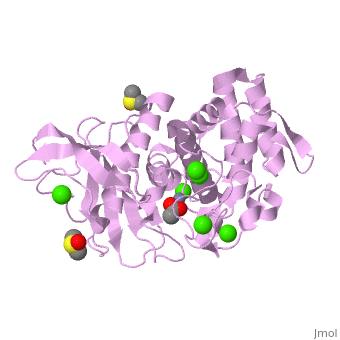This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Thermolysin
From Proteopedia
(Difference between revisions)
| Line 5: | Line 5: | ||
== Structural highlights == | == Structural highlights == | ||
Thermolysin is a well researched [[metalloproteases|metalloprotease]] containing <scene name='User:Ralf_Stephan/Sandbox_2/Zinc/2'>zinc</scene> (click this!) and the amino acids His-Glu-X-His-His as its catalytic center. <scene name='User:Ralf_Stephan/Sandbox_2/Res_yellow/3'>Glu-166, His-142 and -146 are grouped around the zinc atom</scene>, holding it fast, while <scene name='User:Ralf_Stephan/Sandbox_2/Res/1'>Glu-143 holds the polarized water atom. Additionally, Tyr-157 and His-231</scene> stabilize the substrate protein which will be cleaved into two smaller proteins.<ref>Matthews, BW. (1988): ''Structural basis of the action of thermolysin and related zinc peptidases''. In: ''Acc. Chem. Res.'' '''21'''(9); 333–340; http://dx.doi.org/10.1021/ar00153a003</ref><ref>PMID:11935352</ref>. | Thermolysin is a well researched [[metalloproteases|metalloprotease]] containing <scene name='User:Ralf_Stephan/Sandbox_2/Zinc/2'>zinc</scene> (click this!) and the amino acids His-Glu-X-His-His as its catalytic center. <scene name='User:Ralf_Stephan/Sandbox_2/Res_yellow/3'>Glu-166, His-142 and -146 are grouped around the zinc atom</scene>, holding it fast, while <scene name='User:Ralf_Stephan/Sandbox_2/Res/1'>Glu-143 holds the polarized water atom. Additionally, Tyr-157 and His-231</scene> stabilize the substrate protein which will be cleaved into two smaller proteins.<ref>Matthews, BW. (1988): ''Structural basis of the action of thermolysin and related zinc peptidases''. In: ''Acc. Chem. Res.'' '''21'''(9); 333–340; http://dx.doi.org/10.1021/ar00153a003</ref><ref>PMID:11935352</ref>. | ||
| + | |||
| + | == 3D Structures of Thermolysin == | ||
| + | [[Thermolysin 3D structures]] | ||
| + | |||
</StructureSection> | </StructureSection> | ||
| Line 26: | Line 30: | ||
*Thermolysin + amino acid | *Thermolysin + amino acid | ||
| - | **[[1kl6]] – TML+ alanine<br /> | + | **[[1kl6]] – TML residues 1-316 + alanine<br /> |
| - | **[[4m65]] – TML + asparagine<br /> | + | **[[4m65]] – TML residues 1-316 + asparagine<br /> |
**[[8tln]] – TML+ avaline + lysine<br /> | **[[8tln]] – TML+ avaline + lysine<br /> | ||
**[[3qgo]] – TML + phenylalanine methyl ester<br /> | **[[3qgo]] – TML + phenylalanine methyl ester<br /> | ||
| Line 33: | Line 37: | ||
**[[3qh5]] – TML + benzyloxycarboxyl- aspartate + phenylalanine methyl ester | **[[3qh5]] – TML + benzyloxycarboxyl- aspartate + phenylalanine methyl ester | ||
| - | *Thermolysin + | + | *Thermolysin + polypeptide |
| - | **[[2wi0]], [[3tmn]], [[8tln]] – | + | **[[2wi0]], [[3tmn]], [[8tln]] – TMLresidues 1-316 + dipeptide <br /> |
**[[3fgd]] - TML residues 233-548+dipeptide<br /> | **[[3fgd]] - TML residues 233-548+dipeptide<br /> | ||
| + | **[[5onp]], [[5onq]], [[5onr]], [[6ghx]] – TML residues 233-551 + amyloid-β peptide<br /> | ||
*Thermolysin + inhibitor | *Thermolysin + inhibitor | ||
| - | **[[1fjo]], [[1fj3]], [[1fjq]], [[1fjt]], [[1fju]], [[1fjv]], [[1fjw]], [[4tli]], [[5tli]], [[6tli]], [[7tli]], [[8tli]], [[1tli]], [[2tli]], [[3tli]] – TML | + | **[[1fjo]], [[1fj3]], [[1fjq]], [[1fjt]], [[1fju]], [[1fjv]], [[1fjw]], [[4tli]], [[5tli]], [[6tli]], [[7tli]], [[8tli]], [[1tli]], [[2tli]], [[3tli]] – TML residues 1-316 in organic solvents<br /> |
| - | **[[3fv4]], [[3fxp]], [[3flf]], [[3fb0]], [[3fcq]], [[3f28]], [[3f2p]], [[4mtw]], [[4mwp]], [[4mxj]], [[4mzn]], [[4n4e]], [[4n5p]], [[4n66]], [[4oi5]] | + | **[[3fv4]], [[3fxp]], [[3flf]], [[3fb0]], [[3fcq]], [[3f28]], [[3f2p]], [[4mtw]], [[4mwp]], [[4mxj]], [[4mzn]], [[4n4e]], [[4n5p]], [[4n66]], [[4oi5]], [[3for]], [[1y3g]], [[1zdp]], [[1z9g]], [[1os0]], [[1thl]], [[3ms3]], [[3msa]], [[3msf]], [[3msn]], [[3n21]], [[3nn7]], [[3t73]], [[3t74]], [[3t8]], [[3t8c]], [[3t8d]], [[3t8f]], [[3t8g]], [[3t8h]], [[4h57]], [[4d9w]], [[4d91]] – TML residues 233-548+inhibitor |
| - | + | **[[1pe5]], [[6tmn]], [[7tln]], [[4tln]], [[5tln]], [[1hyt]], [[1qf0]], [[1qf1]], [[1qf2]], [[1kjp]], [[1kjo]], [[1kto]], [[1ks7]], [[1kro]], [[1kr6]], [[1kkk]], [[pe7]], [[1pe8]] – TML residues 1-316 + inhibitor <br /> | |
| - | **[[3ssb]] – TML + Impi-α<br /> | + | **[[3ssb]] – TML residues 233-548 + Impi-α<br /> |
| - | **[[3t2h]] – TML + trimethylamine oxide<br /> | + | **[[3t2h]] – TML residues 233-548 + trimethylamine oxide<br /> |
| - | **[[3t2i]] – TML + sarcosine<br /> | + | **[[3t2i]] – TML residues 233-548 + sarcosine<br /> |
| - | **[[3t2j]] – TML + betaine<br /> | + | **[[3t2j]] – TML residues 233-548 + betaine<br /> |
| - | + | **[[6fhp]] – TML C terminal 490-551 + DAIP<br /> | |
| - | **[[6fhp]] – TML C terminal + DAIP<br /> | + | |
| - | + | ||
}} | }} | ||
Revision as of 08:02, 17 February 2020
| |||||||||||
3D Structures of Thermolysin
Updated on 17-February-2020
References
- ↑ Matthews, BW. (1988): Structural basis of the action of thermolysin and related zinc peptidases. In: Acc. Chem. Res. 21(9); 333–340; http://dx.doi.org/10.1021/ar00153a003
- ↑ Pelmenschikov V, Blomberg MR, Siegbahn PE. A theoretical study of the mechanism for peptide hydrolysis by thermolysin. J Biol Inorg Chem. 2002 Mar;7(3):284-98. Epub 2001 Sep 27. PMID:11935352 doi:http://dx.doi.org/10.1007/s007750100295