This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Porphobilinogen synthase
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 1: | Line 1: | ||
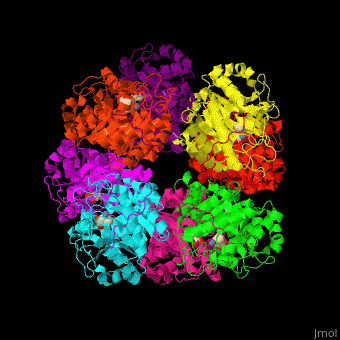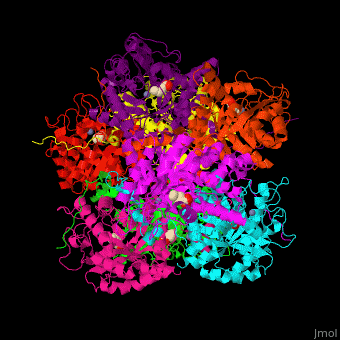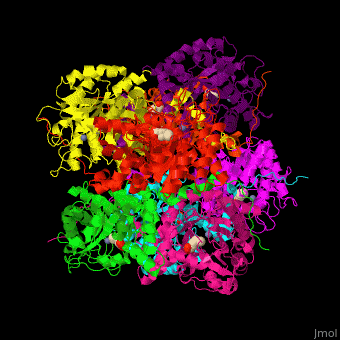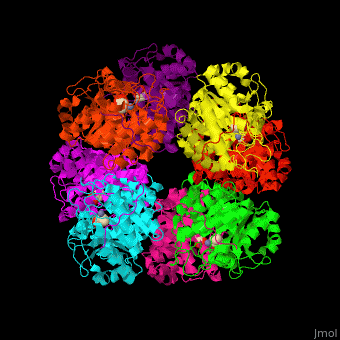
<StructureSection load='' size='450' side='right' caption='Yeast porphobilinogen synthase complex with laevulunic acid and Zn+2 ion (grey) (PDB entry [[1h7n]])' scene='52/526179/Cv/1'> | <StructureSection load='' size='450' side='right' caption='Yeast porphobilinogen synthase complex with laevulunic acid and Zn+2 ion (grey) (PDB entry [[1h7n]])' scene='52/526179/Cv/1'> | ||
== Function == | == Function == | ||
| - | '''Porphobilinogen synthase''' (PBS) catalyzes the formation of porphobilinogen (PBG) from 2 molecules of aminolaevulinic acid (ALA). PBS participates in porphyrin biosynthesis<ref>PMID:15381398</ref>. Levulinic acid (LA) is an inhibitor of PBS. Porphyrin is the precursor of hemes, chlorophyll and vitamin B12. | + | '''Porphobilinogen synthase''' or '''δ-aminolaevulinic acid dehydratase''' (PBS) catalyzes the formation of porphobilinogen (PBG) from 2 molecules of aminolaevulinic acid (ALA). PBS participates in porphyrin biosynthesis<ref>PMID:15381398</ref>. Levulinic acid (LA) is an inhibitor of PBS. Porphyrin is the precursor of hemes, chlorophyll and vitamin B12. |
== Disease == | == Disease == | ||
| Line 17: | Line 17: | ||
*Porphobilinogen synthase | *Porphobilinogen synthase | ||
| + | **[[5hms]], [[5hnr]] – hPBS + Zn – human<br /> | ||
| + | **[[1pv8]] – hPBS (mutant)<br /> | ||
| + | **[[2z0i]], [[2z1b]] - PBS - mouse<br /> | ||
**[[1aw5]] – yPBS (mutant) + Zn – yeast<br /> | **[[1aw5]] – yPBS (mutant) + Zn – yeast<br /> | ||
**[[1qml]] - yPBS + Hg<br /> | **[[1qml]] - yPBS + Hg<br /> | ||
**[[1qnv]] - yPBS + Pb<br /> | **[[1qnv]] - yPBS + Pb<br /> | ||
| - | + | **[[1w54]], [[1w56]], [[1w5m]], [[1w5n]], [[1w5o]], [[1w5p]], [[1w5q]] - PaPBS (mutant) + Mg + Zn – ''Pseudomonas aeruginosa''<br /> | |
| - | + | **[[5lzl]] - PBS + Zn – ''Pyrobaculum calidifontis''<br /> | |
| - | **[[1w54]], [[1w56]], [[1w5m]], [[1w5n]], [[1w5o]], [[1w5p]], [[1w5q]] - PaPBS (mutant) + Mg + Zn – ''Pseudomonas aeruginosa'' | + | |
*Porphobilinogen synthase complex | *Porphobilinogen synthase complex | ||
| - | **[[ | + | **[[1e51]] - hPBS + PBG + Zn<br /> |
| - | + | ||
**[[1ylv]], [[1h7n]] - yPBS + LA + Zn<br /> | **[[1ylv]], [[1h7n]] - yPBS + LA + Zn<br /> | ||
**[[1w31]] - yPBS + LA derivative + Zn<br /> | **[[1w31]] - yPBS + LA derivative + Zn<br /> | ||
| Line 33: | Line 34: | ||
**[[1h7r]], [[1h7p]], [[1eb3]], [[1gjp]] - yPBS + inhibitor + Zn<br /> | **[[1h7r]], [[1h7p]], [[1eb3]], [[1gjp]] - yPBS + inhibitor + Zn<br /> | ||
**[[1ohl]] - yPBS + intermediate + Zn<br /> | **[[1ohl]] - yPBS + intermediate + Zn<br /> | ||
| + | **[[1b4e]] - EcPBS + LA + Zn – ''Escherichia coli''<br /> | ||
| + | **[[5mhb]] - EcPBS + PBG + Zn<br /> | ||
**[[1i8j]], [[1l6s]], [[1l6y]] - EcPBS + inhibitor + Mg + Zn<br /> | **[[1i8j]], [[1l6s]], [[1l6y]] - EcPBS + inhibitor + Mg + Zn<br /> | ||
**[[2c1h]] - PBS + inhibitor + Mg – ''Chlorobium vibrioforme'' <br /> | **[[2c1h]] - PBS + inhibitor + Mg – ''Chlorobium vibrioforme'' <br /> | ||
**[[2c13]], [[2c14]] - PaPBS (mutant) + inhibitor + Mg<br /> | **[[2c13]], [[2c14]] - PaPBS (mutant) + inhibitor + Mg<br /> | ||
**[[2c15]], [[2c16]], [[2c18]], [[2c19]] - PaPBS + inhibitor + Mg<br /> | **[[2c15]], [[2c16]], [[2c18]], [[2c19]] - PaPBS + inhibitor + Mg<br /> | ||
| - | **[[ | + | **[[1b4k]] - PaPBS + LA + Zn<br /> |
**[[1gzg]] - PaPBS + LA derivative + Mg<br /> | **[[1gzg]] - PaPBS + LA derivative + Mg<br /> | ||
**[[2woq]] - PaPBS + antibiotic + Mg<br /> | **[[2woq]] - PaPBS + antibiotic + Mg<br /> | ||
| - | **[[1w1z]] - PBS + LA + Mg – ''Prosthecochloris vibrioformis'' | + | **[[1w1z]] - PBS + LA + Mg – ''Prosthecochloris vibrioformis''<br /> |
| + | **[[3obk]] - PBS residues 303-658 + Mg + PBG – ''Toxoplasma gondii''<br /> | ||
}} | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D structures of porphobilinogen synthase
Updated on 06-October-2020
References
- ↑ Jaffe EK. The porphobilinogen synthase catalyzed reaction mechanism. Bioorg Chem. 2004 Oct;32(5):316-25. PMID:15381398 doi:http://dx.doi.org/10.1016/j.bioorg.2004.05.010
- ↑ Doss M, Muller WA. Acute lead poisoning in inherited porphobilinogen synthase (delta-aminolevulinic acid dehydrase) deficiency. Blut. 1982 Aug;45(2):131-9. PMID:7104498
- ↑ Erskine PT, Newbold R, Brindley AA, Wood SP, Shoolingin-Jordan PM, Warren MJ, Cooper JB. The x-ray structure of yeast 5-aminolaevulinic acid dehydratase complexed with substrate and three inhibitors. J Mol Biol. 2001 Sep 7;312(1):133-41. PMID:11545591 doi:10.1006/jmbi.2001.4947