This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Histone methyltransferase
From Proteopedia
(Difference between revisions)
| (24 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
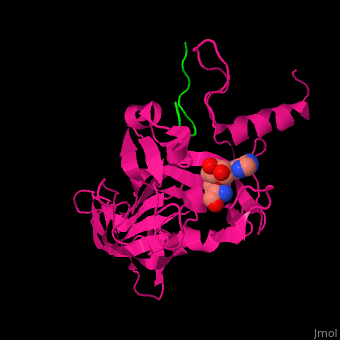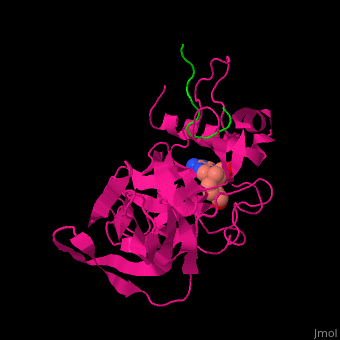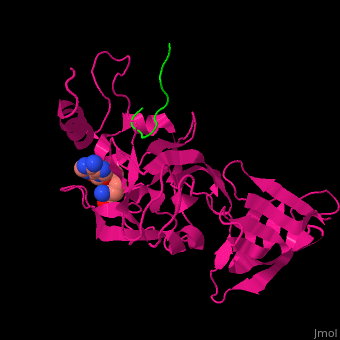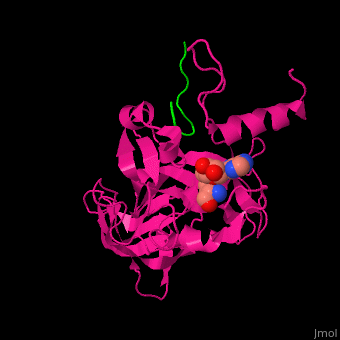
| - | + | <StructureSection load='' size='350' side='right' caption='Human histone-lysine N-methyltrasferase (magenta) complex with methylated lysine histone H3 peptide (green) and S-adenosyl homocysteine (SAH) (PDB entry [[1o9s]])' scene='46/467278/Cv/1'> | |
| - | '''Histone methyltransferase''' (HMT) are histone-lysine N-methyltransferase (KHMT) and histone-arginine N-methyltransferase (RHMT) which catalyzes the transfer of methyl groups to lysine and arginine residues of histones. HMT use S-adenosyl methionine (SAM) or S-adenosyl homocysteine (SAH) as the methyl donor. KHMT can be SET domain-containing or non-SET domain-containing. The SET domain contains the catalytic core of KHMT and targets the lysine tail region of KHMT. | + | == Function == |
| + | '''Histone methyltransferase''' (HMT) are '''histone-lysine N-methyltransferase''' (KHMT) and '''histone-arginine N-methyltransferase''' (RHMT) which catalyzes the transfer of methyl groups to lysine and arginine residues of histones<ref>PMID:15248813</ref>. HMT use S-adenosyl methionine (SAM) or S-adenosyl homocysteine (SAH) as the methyl donor. KHMT can be SET domain-containing or non-SET domain-containing. The SET domain contains the catalytic core of KHMT and targets the lysine tail region of KHMT. CxxC domain is a zinc finger domain. | ||
| - | + | For SET7/9 see [[Histone Lysine Methyltransferase SET7/9]]. | |
| - | == | + | == Relevance == |
| + | Inhibitors of the histone-lysine N-methyltransferase SMYD2 which represses the tumor-suppressing activities of p53 are studied as cancer therapy drugs<ref>PMID:21782458</ref>. | ||
| - | + | == Disease == | |
| + | Aberrant HMT plays a role in cancer, intellectual disability syndromes and regulation of tissue aging<ref>PMID:22473383</ref>. | ||
| - | + | == Structural highlights == | |
| + | Human <scene name='46/467278/Cv/6'>KHMT binds SAH in the peptide binding groove with the histone H3 peptide assuming an extended conformation and the methylated lysine</scene> inserted in a hydrophobic environment<ref>PMID:12540855</ref>. Water molecules shown as red spheres | ||
| - | [[ | + | == 3D Structures of histone methyltransferase == |
| + | [[Histone methyltransferase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | == References == | ||
| + | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Trievel RC. Structure and function of histone methyltransferases. Crit Rev Eukaryot Gene Expr. 2004;14(3):147-69. PMID:15248813
- ↑ Ferguson AD, Larsen NA, Howard T, Pollard H, Green I, Grande C, Cheung T, Garcia-Arenas R, Cowen S, Wu J, Godin R, Chen H, Keen N. Structural Basis of Substrate Methylation and Inhibition of SMYD2. Structure. 2011 Jul 20. PMID:21782458 doi:10.1016/j.str.2011.06.011
- ↑ Greer EL, Shi Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet. 2012 Apr 3;13(5):343-57. doi: 10.1038/nrg3173. PMID:22473383 doi:http://dx.doi.org/10.1038/nrg3173
- ↑ Xiao B, Jing C, Wilson JR, Walker PA, Vasisht N, Kelly G, Howell S, Taylor IA, Blackburn GM, Gamblin SJ. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature. 2003 Feb 6;421(6923):652-6. Epub 2003 Jan 22. PMID:12540855 doi:10.1038/nature01378