This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2exs
From Proteopedia
(Difference between revisions)
(New page: 200px<br /><applet load="2exs" size="450" color="white" frame="true" align="right" spinBox="true" caption="2exs, resolution 2.00Å" /> '''TRAP3 (engineered TR...) |
|||
| (13 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | [[Image:2exs.gif|left|200px]]<br /><applet load="2exs" size="450" color="white" frame="true" align="right" spinBox="true" | ||
| - | caption="2exs, resolution 2.00Å" /> | ||
| - | '''TRAP3 (engineered TRAP)'''<br /> | ||
| - | == | + | ==TRAP3 (engineered TRAP)== |
| - | The protein TRAP (trp RNA binding attenuation protein) forms a highly | + | <StructureSection load='2exs' size='340' side='right'caption='[[2exs]], [[Resolution|resolution]] 2.00Å' scene=''> |
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[2exs]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/Atcc_12980 Atcc 12980]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2EXS OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2EXS FirstGlance]. <br> | ||
| + | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=TRP:TRYPTOPHAN'>TRP</scene></td></tr> | ||
| + | <tr id='NonStdRes'><td class="sblockLbl"><b>[[Non-Standard_Residue|NonStd Res:]]</b></td><td class="sblockDat"><scene name='pdbligand=MSE:SELENOMETHIONINE'>MSE</scene></td></tr> | ||
| + | <tr id='related'><td class="sblockLbl"><b>[[Related_structure|Related:]]</b></td><td class="sblockDat"><div style='overflow: auto; max-height: 3em;'>[[2ext|2ext]]</div></td></tr> | ||
| + | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">mtrB ([https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=1422 ATCC 12980])</td></tr> | ||
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2exs FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2exs OCA], [https://pdbe.org/2exs PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2exs RCSB], [https://www.ebi.ac.uk/pdbsum/2exs PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2exs ProSAT]</span></td></tr> | ||
| + | </table> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/ex/2exs_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=2exs ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
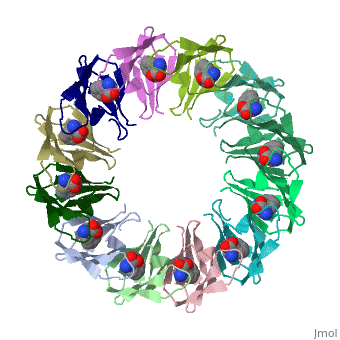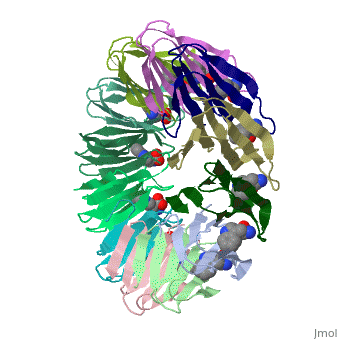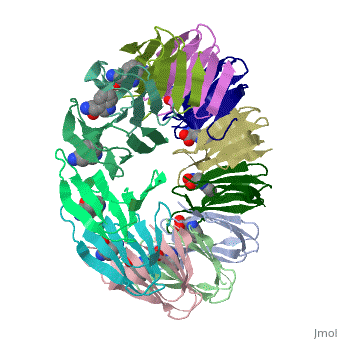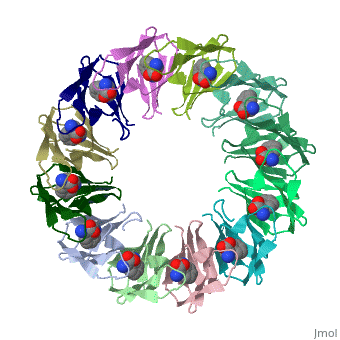
| + | The protein TRAP (trp RNA binding attenuation protein) forms a highly thermostable ring-shaped 11-mer. By linking in tandem two, three, or four DNA sequences encoding TRAP monomers, we have engineered new rings that consist of 12 TRAP subunits and bind 12 ligand molecules. The hydrogen bonding pattern and buried surface area within and between subunits are essentially identical between the 11-mer and 12-mer crystal structures. Why do the artificial proteins choose to make single 12-mer rings? The 12-mer rings are highly sterically strained by their peptide linkers and far from thermostable. That proteins choose to adopt a strained conformation of few subunits rather than an unstrained one with 11 subunits demonstrates the importance of entropic factors in controlling protein-protein interactions in general. | ||
| - | + | Rounding up: Engineering 12-membered rings from the cyclic 11-mer TRAP.,Heddle JG, Yokoyama T, Yamashita I, Park SY, Tame JR Structure. 2006 May;14(5):925-33. PMID:16698553<ref>PMID:16698553</ref> | |
| - | + | ||
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | <div class="pdbe-citations 2exs" style="background-color:#fffaf0;"></div> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ==See Also== | |
| + | *[[Tryptophan RNA-binding attenuation protein|Tryptophan RNA-binding attenuation protein]] | ||
| + | == References == | ||
| + | <references/> | ||
| + | __TOC__ | ||
| + | </StructureSection> | ||
| + | [[Category: Atcc 12980]] | ||
| + | [[Category: Large Structures]] | ||
| + | [[Category: Heddle, J G]] | ||
| + | [[Category: Park, S Y]] | ||
| + | [[Category: Tame, J R.H]] | ||
| + | [[Category: Yamashita, I]] | ||
| + | [[Category: Yokoyama, T]] | ||
| + | [[Category: 12-mer]] | ||
| + | [[Category: Artificial]] | ||
| + | [[Category: Engineered]] | ||
| + | [[Category: Ring protein]] | ||
| + | [[Category: Rna binding protein]] | ||
Current revision
TRAP3 (engineered TRAP)
| |||||||||||