Journal:JBIC:16
From Proteopedia
(Difference between revisions)

| (One intermediate revision not shown.) | |||
| Line 5: | Line 5: | ||
<b>Molecular Tour</b><br> | <b>Molecular Tour</b><br> | ||
| - | Cytochrome c nitrite reductase (ccNIR | + | Cytochrome c nitrite reductase (ccNIR) is a central enzyme of the nitrogen cycle. It binds nitrite, and reduces it by transferring 6 electrons to form ammonia. This ammonia can then be utilized to synthesize nitrogen containing molecules such as amino acids or nucleic acids. However, ccNiR’s primary role is to help extract energy from the reduction; ammonia is simply a potentially useful byproduct. In general, heterotrophic organisms feed on electron-rich substances such as sugars or fatty acids. During the metabolism of these substances large numbers of electrons are produced. Many organisms use oxygen as the final acceptor of these electrons, in which case water is formed. However, some organisms can use alternative electron acceptors such as nitrite, which is where ccNiR comes in. |
The ccNiR described here is produced by the ''Shewanella oneidensis'' bacterium, which is remarkable in its own right due to the large number of electron acceptors that it can utilize. ''Shewanella'' is a facultative anaerobe, which means that it will use oxygen if available, but in the absence of oxygen can get rid of its electrons by dumping them on a wide range of alternate acceptors, of which nitrite is only one example. To handle the electron flow ''Shewanella'' uses a large number of promiscuous <scene name='Journal:JBIC:16/Cv/8'>c-heme</scene> containing electron transfer proteins. Indeed, ''Shewanella'' is exceptionally adept at producing c-heme proteins under fast-growth conditions, which many bacteria commonly used for large-scale laboratory gene expression, such as ''E. coli'', are incapable of unless they are first extensively reprogrammed genetically. Since ''Shewanella'' can be easily grown in the lab, and can naturally and easily produce c-hemes, it is an ideal host for generating large quantities of c-heme proteins such as ccNiR. | The ccNiR described here is produced by the ''Shewanella oneidensis'' bacterium, which is remarkable in its own right due to the large number of electron acceptors that it can utilize. ''Shewanella'' is a facultative anaerobe, which means that it will use oxygen if available, but in the absence of oxygen can get rid of its electrons by dumping them on a wide range of alternate acceptors, of which nitrite is only one example. To handle the electron flow ''Shewanella'' uses a large number of promiscuous <scene name='Journal:JBIC:16/Cv/8'>c-heme</scene> containing electron transfer proteins. Indeed, ''Shewanella'' is exceptionally adept at producing c-heme proteins under fast-growth conditions, which many bacteria commonly used for large-scale laboratory gene expression, such as ''E. coli'', are incapable of unless they are first extensively reprogrammed genetically. Since ''Shewanella'' can be easily grown in the lab, and can naturally and easily produce c-hemes, it is an ideal host for generating large quantities of c-heme proteins such as ccNiR. | ||
The 2012 paper by Youngblut et al. <ref name="Youngblut">none yet</ref> describes a genetically modified ''Shewanella'' strain that can produce 20 – 40 times more ccNiR per liter of culture than the wild type bacterium. The ccNir so produced can be purified easily and in large amounts. This result is important because c-heme proteins have historically proved difficult to over-express in traditional vectors such as ''E. coli''. With large quantities of ''Shewanella'' ccNIR available, Youngblut et al <ref name="Youngblut">none yet</ref> were able to obtain the crystal structure ([[3ubr]]) and do a variety of experiments. The ccNIR consists of <scene name='Journal:JBIC:16/Cv/4'>two equal subunits</scene> (<font color='darkmagenta'><b>colored in darkmagenta</b></font> and <span style="color:lime;background-color:black;font-weight:bold;">in green</span>) with <scene name='Journal:JBIC:16/Cv/5'>five c-hemes each</scene>. In the oxidized ccNIR all central heme irons are Fe3+. They can be subsequently reduced to Fe2+ either by reducing agents or electrochemically. An important conclusion of the paper is that electrons added to ccNiR are likely <scene name='Journal:JBIC:16/Cv/6'>delocalized over several hemes</scene>, rather than localized on individual hemes. | The 2012 paper by Youngblut et al. <ref name="Youngblut">none yet</ref> describes a genetically modified ''Shewanella'' strain that can produce 20 – 40 times more ccNiR per liter of culture than the wild type bacterium. The ccNir so produced can be purified easily and in large amounts. This result is important because c-heme proteins have historically proved difficult to over-express in traditional vectors such as ''E. coli''. With large quantities of ''Shewanella'' ccNIR available, Youngblut et al <ref name="Youngblut">none yet</ref> were able to obtain the crystal structure ([[3ubr]]) and do a variety of experiments. The ccNIR consists of <scene name='Journal:JBIC:16/Cv/4'>two equal subunits</scene> (<font color='darkmagenta'><b>colored in darkmagenta</b></font> and <span style="color:lime;background-color:black;font-weight:bold;">in green</span>) with <scene name='Journal:JBIC:16/Cv/5'>five c-hemes each</scene>. In the oxidized ccNIR all central heme irons are Fe3+. They can be subsequently reduced to Fe2+ either by reducing agents or electrochemically. An important conclusion of the paper is that electrons added to ccNiR are likely <scene name='Journal:JBIC:16/Cv/6'>delocalized over several hemes</scene>, rather than localized on individual hemes. | ||
| Line 12: | Line 12: | ||
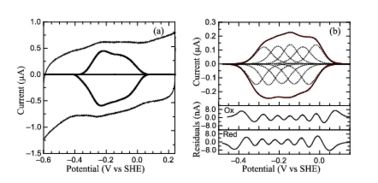
[[Image:figur5.jpg|left|378px|thumb|PFV of ''S. oneidensis'' ccNiR (a) Typical signal on a graphite electrode. (b) Baselinesubtracted non-turnover voltammogram]] | [[Image:figur5.jpg|left|378px|thumb|PFV of ''S. oneidensis'' ccNiR (a) Typical signal on a graphite electrode. (b) Baselinesubtracted non-turnover voltammogram]] | ||
The Ca<sup>2+</sup> ion within <scene name='Journal:JBIC:16/Cv/14'>conserved site</scene> is coordinated in bidentate fashion by <scene name='Journal:JBIC:16/Cv/15'>Glu205</scene>, and in monodentate fashion by the <scene name='Journal:JBIC:16/Cv/16'>Tyr206 and Lys254</scene> backbone carbonyls, and the <scene name='Journal:JBIC:16/Cv/17'>Gln256</scene> side-chain carbonyl. In the ''S. oneidensis'' structure only <scene name='Journal:JBIC:16/Cv/18'>one water molecule</scene> is assigned to the Ca<sup>2+</sup> ion in subunit B. In subunit A the difference electron density that represents this water molecule is very close to the noise level, and it is difficult to identify even one water molecule there. The <scene name='Journal:JBIC:16/Cv/14'>carbonyl side chain of Asp242 and the hydroxyl of Tyr235</scene> come near to the open calcium coordination sites, but are not within bonding distance. Instead they interact with the water molecule that is weakly coordinated to the Ca<sup>2+</sup> ion. The ccNiR calcium ions appear to play a vital role in organizing the <scene name='Journal:JBIC:16/Cv/13'>active site</scene> (as was mentioned above <font color='magenta'><b>hemes-1</b></font> are the active sites). | The Ca<sup>2+</sup> ion within <scene name='Journal:JBIC:16/Cv/14'>conserved site</scene> is coordinated in bidentate fashion by <scene name='Journal:JBIC:16/Cv/15'>Glu205</scene>, and in monodentate fashion by the <scene name='Journal:JBIC:16/Cv/16'>Tyr206 and Lys254</scene> backbone carbonyls, and the <scene name='Journal:JBIC:16/Cv/17'>Gln256</scene> side-chain carbonyl. In the ''S. oneidensis'' structure only <scene name='Journal:JBIC:16/Cv/18'>one water molecule</scene> is assigned to the Ca<sup>2+</sup> ion in subunit B. In subunit A the difference electron density that represents this water molecule is very close to the noise level, and it is difficult to identify even one water molecule there. The <scene name='Journal:JBIC:16/Cv/14'>carbonyl side chain of Asp242 and the hydroxyl of Tyr235</scene> come near to the open calcium coordination sites, but are not within bonding distance. Instead they interact with the water molecule that is weakly coordinated to the Ca<sup>2+</sup> ion. The ccNiR calcium ions appear to play a vital role in organizing the <scene name='Journal:JBIC:16/Cv/13'>active site</scene> (as was mentioned above <font color='magenta'><b>hemes-1</b></font> are the active sites). | ||
| + | |||
| + | '''PDB reference:''' Laue structure of ''Shewanella oneidensis'' cytochrome-c Nitrite Reductase, [[3ubr]]. | ||
</StructureSection> | </StructureSection> | ||
<references/> | <references/> | ||
__NOEDITSECTION__ | __NOEDITSECTION__ | ||
Current revision
| |||||||||||
- ↑ 1.0 1.1 1.2 Youngblut M, Judd ET, Srajer V, Sayyed B, Goelzer T, Elliott SJ, Schmidt M, Pacheco AA. Laue crystal structure of Shewanella oneidensis cytochrome c nitrite reductase from a high-yield expression system. J Biol Inorg Chem. 2012 Mar 2. PMID:22382353 doi:10.1007/s00775-012-0885-0
This page complements a publication in scientific journals and is one of the Proteopedia's Interactive 3D Complement pages. For aditional details please see I3DC.