This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Karsten Theis/overall views
From Proteopedia
(→Introduction) |
|||
| (14 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| + | __NOTOC__ | ||
==Introduction== | ==Introduction== | ||
| + | <gallery> | ||
| + | Image:UvrB fold.JPG | ||
| + | Image:UvrB charge.JPG | ||
| + | Image:UvrB slab core.JPG | ||
| + | Image:UvrB consurf.JPG | ||
| + | </gallery> | ||
This is a collection of how entire protein structures are depicted in publications. The most common views show | This is a collection of how entire protein structures are depicted in publications. The most common views show | ||
* fold of domains | * fold of domains | ||
| Line 8: | Line 15: | ||
==Standard and other views== | ==Standard and other views== | ||
| - | In publications where figures are two dimensional and non-interactive, researchers have to choose a view that shows as much of the interesting features of the protein as possible. Often, when that is not possible, there will be two orthoganal views (e.g. the second rotated by 90 or 180 degrees. The protein used as an example here is the DNA repair enzyme UvrB in complex with ATP (PDB ID 1d9z). This protein not only binds to ATP, but also to DNA and to another DNA repair protein, UvrA. As you look at the various ways protein structures are depicted, you can zoom in to the different binding surfaces or zoom out to the standard view showing the entire protein with the "business" side facing you. | + | In publications where figures are two dimensional and non-interactive, researchers have to choose a view that shows as much of the interesting features of the protein as possible. Often, when that is not possible, there will be two orthoganal views (e.g. the second rotated by 90 or 180 degrees. The protein used as an example here is the DNA repair enzyme UvrB in complex with ATP (PDB ID 1d9z)<ref>PMID:10601012</ref>. This protein not only binds to ATP, but also to DNA and to another DNA repair protein, UvrA. As you look at the various ways protein structures are depicted, you can zoom in to the different binding surfaces or zoom out to the standard view showing the entire protein with the "business" side facing you. |
<jmol> | <jmol> | ||
<jmolButton> | <jmolButton> | ||
| Line 34: | Line 41: | ||
</jmolButton> | </jmolButton> | ||
</jmol> | </jmol> | ||
| + | |||
<jmol> | <jmol> | ||
| Line 42: | Line 50: | ||
</jmolButton> | </jmolButton> | ||
</jmol> | </jmol> | ||
| + | |||
<jmol> | <jmol> | ||
| Line 222: | Line 231: | ||
<script> | <script> | ||
select protein; define ~consurf_to_do selected | select protein; define ~consurf_to_do selected | ||
| - | consurf_initial_scene = true; script "/wiki/ConSurf/d9/1d9z_consurf.spt" | + | consurf_initial_scene = true; script "/wiki/ConSurf/d9/1d9z_consurf.spt";select protein;spacefill only |
</script> | </script> | ||
<text>conservation</text> | <text>conservation</text> | ||
</jmolLink> | </jmolLink> | ||
| - | </jmol>, we use data from [[Introduction to Evolutionary Conservation|ConSurf]]. | + | </jmol>, we use data from [[Introduction to Evolutionary Conservation|ConSurf]] (scene <scene name='78/780454/Conservation/1'>oriented as 2D figure</scene> on the left). |
<jmol> | <jmol> | ||
| Line 285: | Line 294: | ||
<jmol> | <jmol> | ||
<jmolCheckbox> | <jmolCheckbox> | ||
| - | <scriptWhenChecked>select 415-600;rotateselected {558.CA}{580.CA} 120 | + | <scriptWhenChecked>select 415-600;rotateselected {558.CA}{580.CA} 120 </scriptWhenChecked> |
<scriptWhenUnchecked>select 415-600;rotateselected {558.CA}{580.CA} -120</scriptWhenUnchecked> | <scriptWhenUnchecked>select 415-600;rotateselected {558.CA}{580.CA} -120</scriptWhenUnchecked> | ||
<checked>false</checked> | <checked>false</checked> | ||
| Line 384: | Line 393: | ||
| - | This superposition is complicated. Try hiding some of the elements to see clearer, and open a popup 3D browser to view a larger version. Also, rotate the molecules a bit to see them from different angles. You can use the wobble button at the bottom of the page as well. In the original publication [http://emboj.embopress.org/content/18/24/6899.figures-only](Fig. 4), the figure is shown in stereo for better viewing, and domain 2 is omitted. | + | This superposition is complicated. Try hiding some of the elements to see clearer, and open a popup 3D browser to view a larger version. Also, rotate the molecules a bit to see them from different angles. You can use the wobble button at the bottom of the page as well. In the original publication [http://emboj.embopress.org/content/18/24/6899.figures-only](Fig. 4), the figure is shown in stereo for better viewing, and domain 2 is omitted. In the Jmol browser used here, you can turn on stereo as well. This is best done in the pop-up window using the right-click menu, but depending on your eyes, you might need stereo glasses to experience the effect. |
<jmol> | <jmol> | ||
Current revision
Introduction
This is a collection of how entire protein structures are depicted in publications. The most common views show
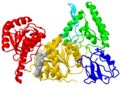
- fold of domains
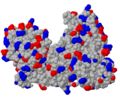
- charge distribution
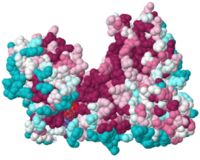
- hydrophobic patches
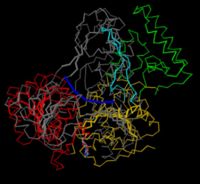
- surface conservation
- superpositions with related structures
Standard and other views
In publications where figures are two dimensional and non-interactive, researchers have to choose a view that shows as much of the interesting features of the protein as possible. Often, when that is not possible, there will be two orthoganal views (e.g. the second rotated by 90 or 180 degrees. The protein used as an example here is the DNA repair enzyme UvrB in complex with ATP (PDB ID 1d9z)[1]. This protein not only binds to ATP, but also to DNA and to another DNA repair protein, UvrA. As you look at the various ways protein structures are depicted, you can zoom in to the different binding surfaces or zoom out to the standard view showing the entire protein with the "business" side facing you.
Types of overall views
| |||||||||||
References
- ↑ Theis K, Chen PJ, Skorvaga M, Van Houten B, Kisker C. Crystal structure of UvrB, a DNA helicase adapted for nucleotide excision repair. EMBO J. 1999 Dec 15;18(24):6899-907. PMID:10601012 doi:10.1093/emboj/18.24.6899