This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Aconitase
From Proteopedia
(Difference between revisions)
| (5 intermediate revisions not shown.) | |||
| Line 3: | Line 3: | ||
==Function== | ==Function== | ||
| - | [[Aconitase]] (ACO, EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=4.2.1.3 4.2.1.3]) is an enzymatic domain that confers the ability to catalyse the equilibrium | + | [[Aconitase]] (ACO, EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=4.2.1.3 4.2.1.3]) is an enzymatic domain that confers the ability to catalyse the equilibrium |
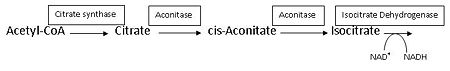
:citrate = aconitate + H<sub>2</sub>O = L-isocitrate | :citrate = aconitate + H<sub>2</sub>O = L-isocitrate | ||
This reaction is part of the citrate (TCA-, Krebs-)cycle. | This reaction is part of the citrate (TCA-, Krebs-)cycle. | ||
In most organisms, there is a cytosolic enzyme with an ACO domain (cAc), and in eukaryotes, a second copy of it was introduced with mitochondria (mAc). Plants developed even more copies in mitochondria. | In most organisms, there is a cytosolic enzyme with an ACO domain (cAc), and in eukaryotes, a second copy of it was introduced with mitochondria (mAc). Plants developed even more copies in mitochondria. | ||
| - | Aconitase contains a Fe4S4 cluster which converts to Fe3S4 when the enzyme is inactive. In humans, two types of ACO are expressed: the soluble '''ACO1''' and the mitochondrial '''ACO2'''. | + | Aconitase contains a Fe4S4 cluster which converts to Fe3S4 when the enzyme is inactive. In humans, two types of ACO are expressed: the soluble '''ACO1''' and the mitochondrial '''ACO2'''. Two types of '''ACO X''' were characterized as '''mevalonate 5-phosphate dehydratase''' and '''cis-3-hydroxy-L-proline dehydrates'''. |
Aconitase from pig (PDB [[7acn]]) is a single polypeptide (M<sub>r</sub> 83kD) that catalyzes the reversible isomerization of citrate and isocitrate.<ref name="Zheng">PMID 1313811</ref> It is the second enzyme in the Citric acid cycle, which is a series of enzyme-catalysed chemical reactions that is crucial to aerobic cellular respiration and the production of ATP. See also:<br /> | Aconitase from pig (PDB [[7acn]]) is a single polypeptide (M<sub>r</sub> 83kD) that catalyzes the reversible isomerization of citrate and isocitrate.<ref name="Zheng">PMID 1313811</ref> It is the second enzyme in the Citric acid cycle, which is a series of enzyme-catalysed chemical reactions that is crucial to aerobic cellular respiration and the production of ATP. See also:<br /> | ||
*[[Citric Acid Cycle]] | *[[Citric Acid Cycle]] | ||
*[[Krebs cycle step 2]] | *[[Krebs cycle step 2]] | ||
| + | *[[Glyoxylate cycle]] | ||
==Structure== | ==Structure== | ||
| Line 20: | Line 21: | ||
== Catalytic mechanism of mitochondrial ACO == | == Catalytic mechanism of mitochondrial ACO == | ||
| - | Both mAc and cAc are quite similar in their ACO function. Studies, however, concentrated on <scene name='Aconitase/7acn-sf4/1'>the mitochondrial ACO</scene>. ACO is an excellent system for understanding the role of iron-sulfur-clusters in catalysis. The <scene name='Aconitase/7acn-sf4/2'>(4Fe-4S) cofactor is held in place</scene> by three sulfur atoms belonging to the cysteins-385, -448, and -451 <scene name=' | + | Both mAc and cAc are quite similar in their ACO function. Studies, however, concentrated on <scene name='Aconitase/7acn-sf4/1'>the mitochondrial ACO</scene>. ACO is an excellent system for understanding the role of iron-sulfur-clusters in catalysis. The <scene name='Aconitase/7acn-sf4/2'>(4Fe-4S) cofactor is held in place</scene> by three sulfur atoms belonging to the cysteins-385, -448, and -451 <scene name='33/338089/7acn-morph/5'>which are bound to three of the four</scene> cluster iron atoms. On activation of the enzyme, <scene name='33/338089/7acn-morph/8'>a fourth iron atom is included in the cluster</scene> together with a water molecule.This Fe4 is free to bind one, two, or three partners, in this reaction always oxygen atoms belonging to other molecules.<ref>PMID:8151704</ref> |
<!--It is clear that, in order to synthesize L-isocitrate, stereoselective catalysis must occur.--> | <!--It is clear that, in order to synthesize L-isocitrate, stereoselective catalysis must occur.--> | ||
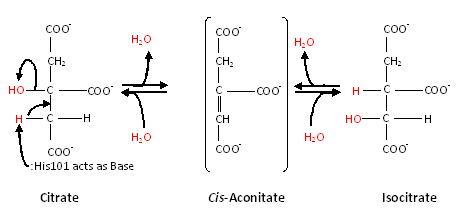
Substrate-free aconitase contains a [4Fe-4S]<sup>2+</sup> cluster with hydroxyl bound to one of the Fe. Upon binding of substrate the bound hydroxyl is protonated. A hydrogen bond from <scene name='Anthony_Noles_Sandbox/His101/3'>His101</scene> to the isocitrate hydroxyl is donated to form water. Alternatively, the proton could be donated by <scene name='Anthony_Noles_Sandbox/His167/3'>His167</scene> as this histidine is hydrogen bonded to a H<sub>2</sub>O molecule. His167 is also hydrogen bonded to the bound H<sub>2</sub>O in the [4Fe-4S] cluster. Both <scene name='Anthony_Noles_Sandbox/His_101_and_167/4'>His101 and His167</scene> are paired with carboxylates (<scene name='Anthony_Noles_Sandbox/Asp100_and_glu262/3'>Asp100 and Glu262</scene>, respectively) and are likely to be protonated. The conformational change associated with substrate binding reorients the cluster. <ref name="Beinert" /> The residue which removes a proton from citrate or isocitrate is <scene name='Anthony_Noles_Sandbox/Ser642/4'>Ser642</scene>. <ref name="Beinert" /> This causes the cis-Aconitate intermediate (seen below), which consists of a double bond, which is a direct result of the deprotonation. Then, there is a rehydration of the double bond of cis-aconitate to form isocitrate (if the original substrate was citrate). To better understand this, consider this process as stages, seen below. | Substrate-free aconitase contains a [4Fe-4S]<sup>2+</sup> cluster with hydroxyl bound to one of the Fe. Upon binding of substrate the bound hydroxyl is protonated. A hydrogen bond from <scene name='Anthony_Noles_Sandbox/His101/3'>His101</scene> to the isocitrate hydroxyl is donated to form water. Alternatively, the proton could be donated by <scene name='Anthony_Noles_Sandbox/His167/3'>His167</scene> as this histidine is hydrogen bonded to a H<sub>2</sub>O molecule. His167 is also hydrogen bonded to the bound H<sub>2</sub>O in the [4Fe-4S] cluster. Both <scene name='Anthony_Noles_Sandbox/His_101_and_167/4'>His101 and His167</scene> are paired with carboxylates (<scene name='Anthony_Noles_Sandbox/Asp100_and_glu262/3'>Asp100 and Glu262</scene>, respectively) and are likely to be protonated. The conformational change associated with substrate binding reorients the cluster. <ref name="Beinert" /> The residue which removes a proton from citrate or isocitrate is <scene name='Anthony_Noles_Sandbox/Ser642/4'>Ser642</scene>. <ref name="Beinert" /> This causes the cis-Aconitate intermediate (seen below), which consists of a double bond, which is a direct result of the deprotonation. Then, there is a rehydration of the double bond of cis-aconitate to form isocitrate (if the original substrate was citrate). To better understand this, consider this process as stages, seen below. | ||
| Line 52: | Line 53: | ||
</StructureSection> | </StructureSection> | ||
__NOTOC__ | __NOTOC__ | ||
| - | == 3D structures of Aconitase== | ||
| - | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| - | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
| - | |||
| - | * ACO | ||
| - | |||
| - | **[[1b0k]] – pACO (mutant) – pig<br /> | ||
| - | **[[5acn]] – pACO+Fe3S4<br /> | ||
| - | **[[6acn]] - pACO+Fe4S4<br /> | ||
| - | **[[1amj]], [[1nit]] – cACO - cow<br /> | ||
| - | |||
| - | *ACO+citrate | ||
| - | |||
| - | **[[1c96]] - pACO (mutant)+citrate<br /> | ||
| - | **[[1b0m]] - pACO (mutant)+fluorocitrate<br /> | ||
| - | |||
| - | *ACO+aconitate | ||
| - | |||
| - | **[[1fgh]] – cACO+4-hydroxy-aconitate <br /> | ||
| - | **[[1aco]] – cACO+transaconitate<br /> | ||
| - | **[[1nis]] - cACO+transaconitate+nitrocitrate<br /> | ||
| - | |||
| - | *ACO+isocitrate | ||
| - | |||
| - | **[[7acn]] - pACO +isocitrate<br /> | ||
| - | **[[1c97]], [[1b0j]] - pACO (mutant)+isocitrate<br /> | ||
| - | **[[1ami]], [[8acn]] – cACO+isocitrate<br /> | ||
| - | |||
| - | * ACO1 | ||
| - | |||
| - | **[[2b3x]], [[2b3y]] – hACO1 – human<br /> | ||
| - | **[[3snp]] – rACO1 (mutant)+ferritin H IRE-RNA – rabbit<br /> | ||
| - | **[[3sn2]] - rACO1 (mutant)+ transferrin receptor iron regulatory RNA<br /> | ||
| - | |||
| - | * ACO2 | ||
| - | **[[1l5j]] – ACO2 – ''Escherichia coli''<br /> | ||
| - | }} | ||
== Literature == | == Literature == | ||
Current revision
| |||||||||||
Literature
- M. Claire Kennedy and Helmut Beinert: IX.4. Aconitase. in Ivano Bertini, Harry B. Gray, Edward I. Stiefel, Joan Selverstone Valentine (eds.): Biological Inorganic Chemistry: Structure and Reactivity. University Science Books, Herndon 2006. ISBN 1891389432 pp.209--
Additional Resources
For additional information, see: Carbohydrate Metabolism; Krebs cycle step 2.
References
- ↑ Zheng L, Kennedy MC, Beinert H, Zalkin H. Mutational analysis of active site residues in pig heart aconitase. J Biol Chem. 1992 Apr 15;267(11):7895-903. PMID:1313811
- ↑ 2.0 2.1 Frishman D, Hentze MW. Conservation of aconitase residues revealed by multiple sequence analysis. Implications for structure/function relationships. Eur J Biochem. 1996 Jul 1;239(1):197-200. PMID:8706708
- ↑ Dupuy J, Volbeda A, Carpentier P, Darnault C, Moulis JM, Fontecilla-Camps JC. Crystal structure of human iron regulatory protein 1 as cytosolic aconitase. Structure. 2006 Jan;14(1):129-39. PMID:16407072 doi:10.1016/j.str.2005.09.009
- ↑ 4.0 4.1 4.2 Beinert, H., Kennedy, M. C., Stout, C.D. “Aconitase as Iron−Sulfur Protein, Enzyme, and Iron-Regulatory Protein.” Chem. Rev. 1996, 96, 2335−2373.
- ↑ Lauble H, Kennedy MC, Beinert H, Stout CD. Crystal structures of aconitase with trans-aconitate and nitrocitrate bound. J Mol Biol. 1994 Apr 8;237(4):437-51. PMID:8151704 doi:http://dx.doi.org/10.1006/jmbi.1994.1246
- ↑ 6.0 6.1 6.2 6.3 Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. p. 578-579. Print.
- ↑ 7.0 7.1 Flint, DH., and Allen, RM. "Iron-sulfur protein with nonredox functions.” Chem. Rev. 1996, 96, 2315−2334.
External links
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Ralf Stephan, David Canner, Joel L. Sussman, Jaime Prilusky, Anthony Noles, Angel Herraez, Eran Hodis