This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Malate synthase
From Proteopedia
(Difference between revisions)
| (6 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
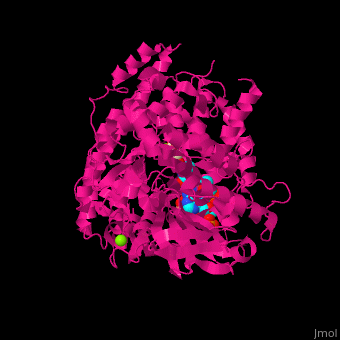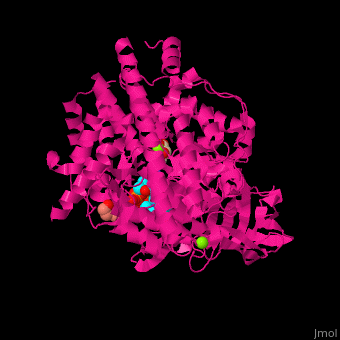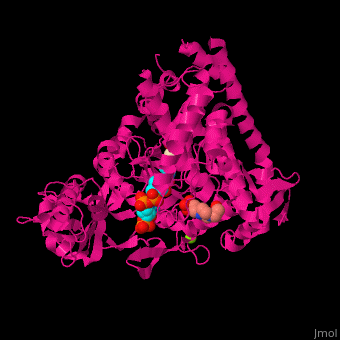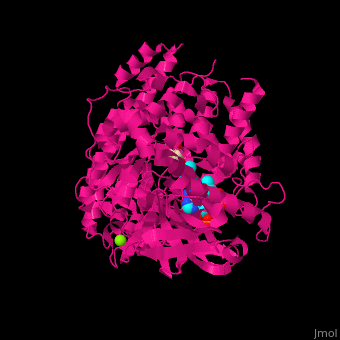
| - | <StructureSection load=' | + | <StructureSection load='' size='350' side='right' caption='Structure of malate synthase G complex with CoA, malate, Hepes and Mg+2 ion (green) (PDB entry [[2gq3]])' scene='57/573146/Cv/1'> |
== Function == | == Function == | ||
| - | '''Malate synthase''' (MS) catalyzes the reversible conversion of acetyl-CoA, glyoxylate and water to (S)-malate and CoA. MS participates in pyruvate, glyoxylate and dicarboxylate metabolism. There are 2 isozymes of MS. '''MS G''' is formed during growth on glycolate<ref>PMID:12930982</ref> and '''MS A''' which metabolizes glyoxylate formed in the dissimilation of acetate. | + | '''Malate synthase''' (MS) catalyzes the reversible conversion of acetyl-CoA, glyoxylate and water to [[R/S nomenclature|(S)]]-malate and CoA. MS participates in pyruvate, glyoxylate and dicarboxylate metabolism. There are 2 isozymes of MS. '''MS G''' is formed during growth on glycolate<ref>PMID:12930982</ref> and '''MS A''' which metabolizes glyoxylate formed in the dissimilation of acetate. |
| + | |||
| + | See also: [[Glyoxylate cycle]] | ||
== Structural highlights == | == Structural highlights == | ||
| - | <scene name='57/573146/Cv/ | + | <scene name='57/573146/Cv/8'>MS active site pocket is situated between the TIM barrel and the C-terminal</scene>. The ternary complex contains malate, acetyl-CoA and Mg+2 ion<ref>PMID:16877713</ref>. |
| - | <scene name='57/573146/Cv/ | + | <scene name='57/573146/Cv/9'>Malate/Mg+2 ion binding site</scene>. Water molecules are shown as red spheres. |
| - | <scene name='57/573146/Cv/ | + | <scene name='57/573146/Cv/10'>Acetyl-CoA binding site</scene>. |
| - | + | ||
==3d structures of malate synthase== | ==3d structures of malate synthase== | ||
| + | [[Malate synthase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | **[[1p7t]] – EcMS G (mutant) + pyruvate + Mg + acetyl-CoA<br /> | ||
| - | **[[3oyz]] – HvMS + pyruvate + Mg + acetyl-CoA<br /> | ||
| - | **[[1n8w]] – MtMS G + glyoxylate + Mg + CoA<br /> | ||
| - | **[[2gq3]] – MtMS G + malate + Mg + CoA<br /> | ||
| - | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Anstrom DM, Kallio K, Remington SJ. Structure of the Escherichia coli malate synthase G:pyruvate:acetyl-coenzyme A abortive ternary complex at 1.95 A resolution. Protein Sci. 2003 Sep;12(9):1822-32. PMID:12930982
- ↑ Anstrom DM, Remington SJ. The product complex of M. tuberculosis malate synthase revisited. Protein Sci. 2006 Aug;15(8):2002-7. PMID:16877713 doi:15/8/2002
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Karsten Theis, Joel L. Sussman