This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
3hts
From Proteopedia
(Difference between revisions)
| (10 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | {{Seed}} | ||
| - | [[Image:3hts.png|left|200px]] | ||
| - | < | + | ==HEAT SHOCK TRANSCRIPTION FACTOR/DNA COMPLEX== |
| - | + | <StructureSection load='3hts' size='340' side='right'caption='[[3hts]], [[Resolution|resolution]] 1.75Å' scene=''> | |
| - | You may | + | == Structural highlights == |
| - | + | <table><tr><td colspan='2'>[[3hts]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/Kluyveromyces_lactis Kluyveromyces lactis]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=3HTS OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=3HTS FirstGlance]. <br> | |
| - | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 1.75Å</td></tr> | |
| - | - | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=GOL:GLYCEROL'>GOL</scene></td></tr> |
| - | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=3hts FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3hts OCA], [https://pdbe.org/3hts PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=3hts RCSB], [https://www.ebi.ac.uk/pdbsum/3hts PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=3hts ProSAT]</span></td></tr> | |
| + | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/HSF_KLULA HSF_KLULA] DNA-binding protein that specifically binds heat shock promoter elements (HSE) and activates transcription. Also required for growth at normal temperatures. | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/ht/3hts_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=3hts ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
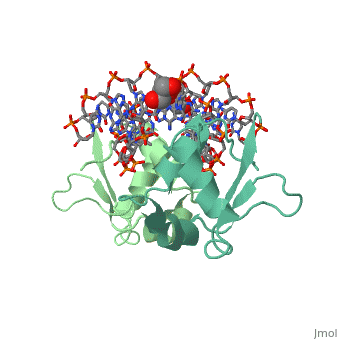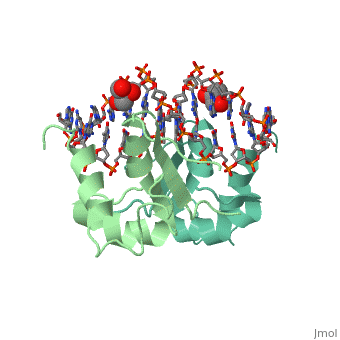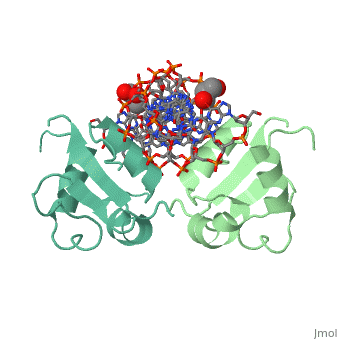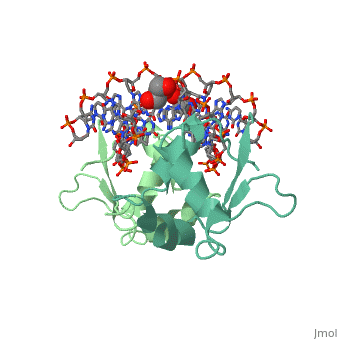
| + | The 1.75 A crystal structure of the Kluyveromyces lactis heat shock transcription factor (HSF) DNA-binding domain (DBD) complexed with DNA reveals a protein-DNA interface with few direct major groove contacts and a number of phosphate backbone contacts that are primarily water-mediated interactions. The DBD, a 'winged' helix-turn-helix protein, displays a novel mode of binding in that the 'wing' does not contact DNA like all others of that class. Instead, the monomeric DBD, which crystallized as a symmetric dimer to a pair of nGAAn inverted repeats, uses the 'wing' to form part of the protein-protein contacts. This dimer interface is likely important for increasing the DNA-binding specificity and affinity of the trimeric form of HSF, as well as for increasing cooperativity between adjacent trimers. | ||
| - | + | A new use for the 'wing' of the 'winged' helix-turn-helix motif in the HSF-DNA cocrystal.,Littlefield O, Nelson HC Nat Struct Biol. 1999 May;6(5):464-70. PMID:10331875<ref>PMID:10331875</ref> | |
| + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| + | </div> | ||
| + | <div class="pdbe-citations 3hts" style="background-color:#fffaf0;"></div> | ||
| - | + | ==See Also== | |
| - | + | *[[Heat shock factor|Heat shock factor]] | |
| - | + | == References == | |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | + | </StructureSection> | |
| - | == | + | |
| - | + | ||
| - | + | ||
| - | == | + | |
| - | < | + | |
[[Category: Kluyveromyces lactis]] | [[Category: Kluyveromyces lactis]] | ||
| - | [[Category: | + | [[Category: Large Structures]] |
| - | [[Category: | + | [[Category: Littlefield O]] |
| - | [[Category: | + | [[Category: Nelson HCM]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
HEAT SHOCK TRANSCRIPTION FACTOR/DNA COMPLEX
| |||||||||||