This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2cfp
From Proteopedia
(Difference between revisions)
| (6 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | {{STRUCTURE_2cfp| PDB=2cfp | SCENE= }} | ||
| - | ===SUGAR FREE LACTOSE PERMEASE AT ACIDIC PH=== | ||
| - | {{ABSTRACT_PUBMED_16525509}} | ||
| - | == | + | ==Sugar Free Lactose Permease at acidic pH== |
| - | [[2cfp]] is a 1 chain structure with sequence from [ | + | <StructureSection load='2cfp' size='340' side='right'caption='[[2cfp]], [[Resolution|resolution]] 3.30Å' scene=''> |
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[2cfp]] is a 1 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_coli Escherichia coli]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2CFP OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2CFP FirstGlance]. <br> | ||
| + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 3.3Å</td></tr> | ||
| + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=HG:MERCURY+(II)+ION'>HG</scene></td></tr> | ||
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2cfp FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2cfp OCA], [https://pdbe.org/2cfp PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2cfp RCSB], [https://www.ebi.ac.uk/pdbsum/2cfp PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2cfp ProSAT]</span></td></tr> | ||
| + | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/LACY_ECOLI LACY_ECOLI] Responsible for transport of beta-galactosides into the cell, with the concomitant import of a proton (symport system). | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/cf/2cfp_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=2cfp ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
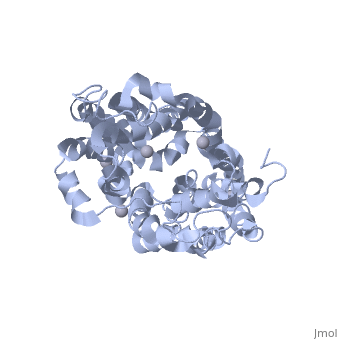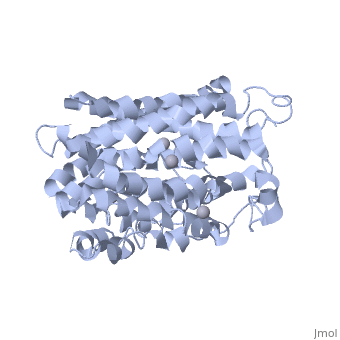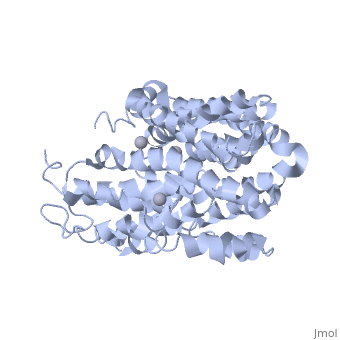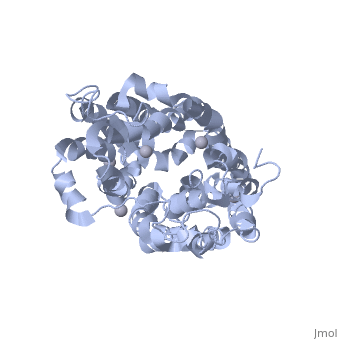
| + | Cation-coupled active transport is an essential cellular process found ubiquitously in all living organisms. Here, we present two novel ligand-free X-ray structures of the lactose permease (LacY) of Escherichia coli determined at acidic and neutral pH, and propose a model for the mechanism of coupling between lactose and H+ translocation. No sugar-binding site is observed in the absence of ligand, and deprotonation of the key residue Glu269 is associated with ligand binding. Thus, substrate induces formation of the sugar-binding site, as well as the initial step in H+ transduction. | ||
| + | |||
| + | Structural evidence for induced fit and a mechanism for sugar/H+ symport in LacY.,Mirza O, Guan L, Verner G, Iwata S, Kaback HR EMBO J. 2006 Mar 22;25(6):1177-83. Epub 2006 Mar 9. PMID:16525509<ref>PMID:16525509</ref> | ||
| + | |||
| + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| + | </div> | ||
| + | <div class="pdbe-citations 2cfp" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
*[[Lactose Permease|Lactose Permease]] | *[[Lactose Permease|Lactose Permease]] | ||
| - | + | == References == | |
| - | + | <references/> | |
| - | == | + | __TOC__ |
| - | < | + | </StructureSection> |
[[Category: Escherichia coli]] | [[Category: Escherichia coli]] | ||
| - | [[Category: Guan | + | [[Category: Large Structures]] |
| - | [[Category: Iwata | + | [[Category: Guan L]] |
| - | [[Category: Kaback | + | [[Category: Iwata S]] |
| - | [[Category: Mirza | + | [[Category: Kaback HR]] |
| - | [[Category: Verner | + | [[Category: Mirza O]] |
| - | + | [[Category: Verner G]] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
Sugar Free Lactose Permease at acidic pH
| |||||||||||
Categories: Escherichia coli | Large Structures | Guan L | Iwata S | Kaback HR | Mirza O | Verner G