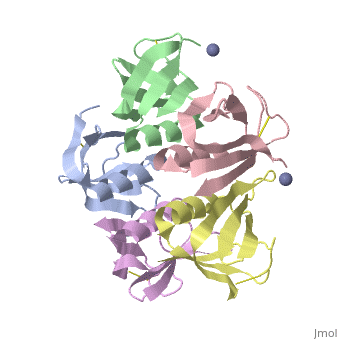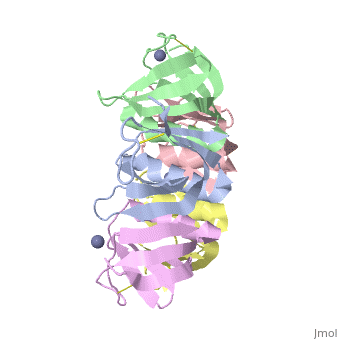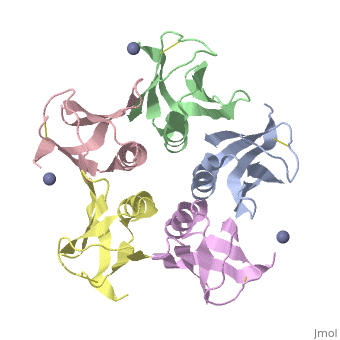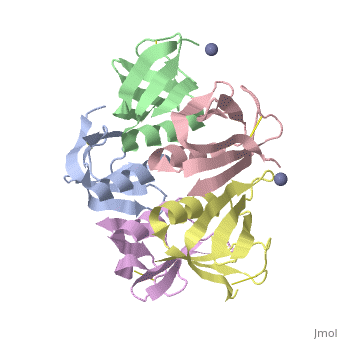This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2xsc
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Structural highlights == | == Structural highlights == | ||
<table><tr><td colspan='2'>[[2xsc]] is a 5 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_coli Escherichia coli]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1bov 1bov]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2XSC OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2XSC FirstGlance]. <br> | <table><tr><td colspan='2'>[[2xsc]] is a 5 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_coli Escherichia coli]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1bov 1bov]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2XSC OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2XSC FirstGlance]. <br> | ||
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.052Å</td></tr> |
| - | <tr id=' | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=ZN:ZINC+ION'>ZN</scene></td></tr> |
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2xsc FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2xsc OCA], [https://pdbe.org/2xsc PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2xsc RCSB], [https://www.ebi.ac.uk/pdbsum/2xsc PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2xsc ProSAT]</span></td></tr> | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2xsc FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2xsc OCA], [https://pdbe.org/2xsc PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2xsc RCSB], [https://www.ebi.ac.uk/pdbsum/2xsc PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2xsc ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
| - | + | [https://www.uniprot.org/uniprot/STXB_BPH19 STXB_BPH19] The B subunit is responsible for the binding of the holotoxin to specific receptors on the target cell surface, such as globotriaosylceramide (Gb3) in human intestinal microvilli. | |
<div style="background-color:#fffaf0;"> | <div style="background-color:#fffaf0;"> | ||
== Publication Abstract from PubMed == | == Publication Abstract from PubMed == | ||
| Line 29: | Line 29: | ||
[[Category: Escherichia coli]] | [[Category: Escherichia coli]] | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: Boodhoo | + | [[Category: Boodhoo A]] |
| - | [[Category: Brunton | + | [[Category: Brunton JL]] |
| - | [[Category: Bunkoczi | + | [[Category: Bunkoczi G]] |
| - | [[Category: Oeffner | + | [[Category: Oeffner RD]] |
| - | [[Category: Read | + | [[Category: Read RJ]] |
| - | [[Category: Stein | + | [[Category: Stein PE]] |
| - | [[Category: Tyrrell | + | [[Category: Tyrrell GJ]] |
| - | + | ||
Current revision
Crystal structure of the cell-binding B oligomer of verotoxin-1 from E. coli
| |||||||||||