This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1xbr
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='1xbr' size='340' side='right'caption='[[1xbr]], [[Resolution|resolution]] 2.50Å' scene=''> | <StructureSection load='1xbr' size='340' side='right'caption='[[1xbr]], [[Resolution|resolution]] 2.50Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1xbr]] is a 4 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1xbr]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Xenopus_laevis Xenopus laevis]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1XBR OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1XBR FirstGlance]. <br> |
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.5Å</td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1xbr FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1xbr OCA], [https://pdbe.org/1xbr PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1xbr RCSB], [https://www.ebi.ac.uk/pdbsum/1xbr PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1xbr ProSAT]</span></td></tr> |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [ | + | [https://www.uniprot.org/uniprot/TBXT_XENLA TBXT_XENLA] Involved in the transcriptional regulation of genes required for mesoderm formation and differentiation (PubMed:11869292). Binds to the palindromic T site 5'-TTCACACCTAGGTGTGAA-3' DNA sequence (PubMed:11111039). Causes dorsal mesodermal differentiation of animal cap ectoderm when co-expressed with wnt8 and noggin (PubMed:7906224). None of these molecules causes dorsal mesoderm formation when expressed alone (PubMed:7906224). Establishes the left/right axis at early gastrula stage by directly up-regulating mesodermal expression of zic3 (PubMed:11869292).<ref>PMID:11111039</ref> <ref>PMID:11869292</ref> <ref>PMID:7906224</ref> |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 19: | Line 19: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1xbr ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1xbr ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
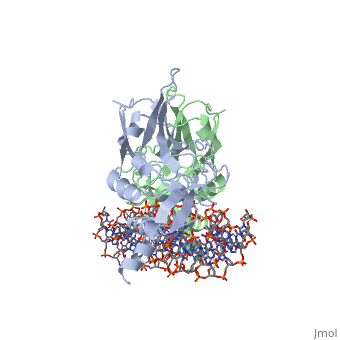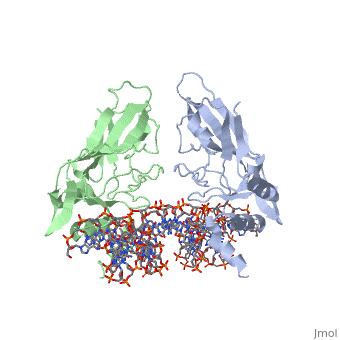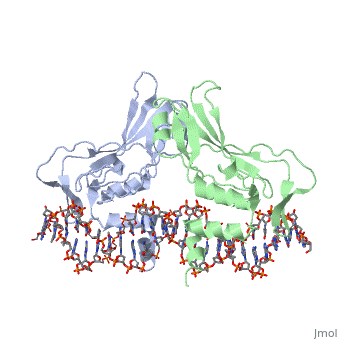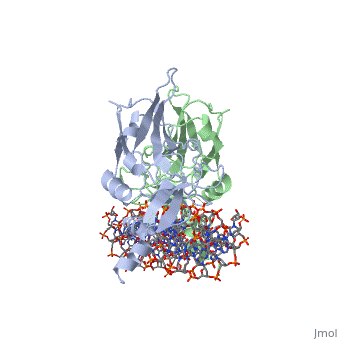
| - | The mouse Brachyury (T) gene is the prototype of a growing family of so-called T-box genes which encode transcriptional regulators and have been identified in a variety of invertebrates and vertebrates, including humans. Mutations in Brachyury and other T-box genes result in drastic embryonic phenotypes, indicating that T-box gene products are essential in tissue specification, morphogenesis and organogenesis. The T-box encodes a DNA-binding domain of about 180 amino-acid residues, the T domain. Here we report the X-ray structure of the T domain from Xenopus laevis in complex with a 24-nucleotide palindromic DNA duplex. We show that the protein is bound as a dimer, interacting with the major and the minor grooves of the DNA. A new type of specific DNA contact is seen, in which a carboxy-terminal helix is deeply embedded into an enlarged minor groove without bending the DNA. Hydrophobic interactions and an unusual main-chain carbonyl contact to a guanine account for sequence-specific recognition in the minor groove by this helix. Thus the structure of this T domain complex with DNA reveals a new way in which a protein can recognize DNA. | ||
| - | |||
| - | Crystallographic structure of the T domain-DNA complex of the Brachyury transcription factor.,Muller CW, Herrmann BG Nature. 1997 Oct 23;389(6653):884-8. PMID:9349824<ref>PMID:9349824</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 1xbr" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
| Line 35: | Line 26: | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
| - | [[Category: African clawed frog]] | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: | + | [[Category: Xenopus laevis]] |
| - | [[Category: | + | [[Category: Muller CW]] |
| - | + | ||
| - | + | ||
Current revision
T DOMAIN FROM XENOPUS LAEVIS BOUND TO DNA
| |||||||||||