This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
3kyl
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='3kyl' size='340' side='right'caption='[[3kyl]], [[Resolution|resolution]] 2.70Å' scene=''> | <StructureSection load='3kyl' size='340' side='right'caption='[[3kyl]], [[Resolution|resolution]] 2.70Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[3kyl]] is a 2 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[3kyl]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/Tribolium_castaneum Tribolium castaneum]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=3KYL OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=3KYL FirstGlance]. <br> |
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.7Å</td></tr> |
| - | <tr id=' | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=MG:MAGNESIUM+ION'>MG</scene></td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=3kyl FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3kyl OCA], [https://pdbe.org/3kyl PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=3kyl RCSB], [https://www.ebi.ac.uk/pdbsum/3kyl PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=3kyl ProSAT]</span></td></tr> |
</table> | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/Q0QHL8_TRICA Q0QHL8_TRICA] | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 18: | Line 20: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=3kyl ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=3kyl ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
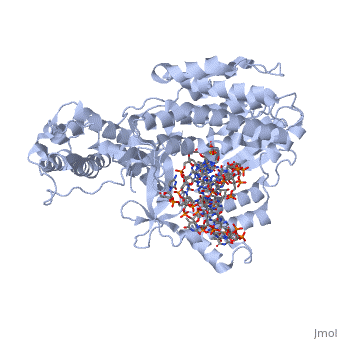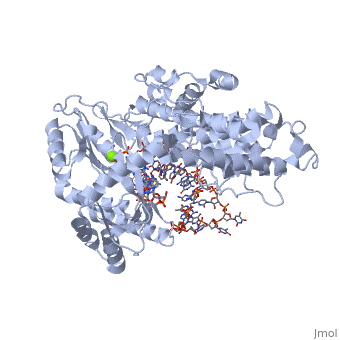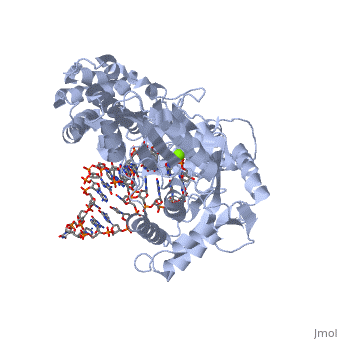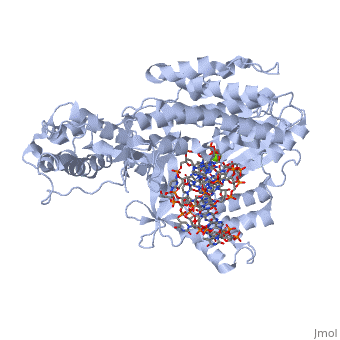
| - | Telomerase is a specialized DNA polymerase that extends the 3' ends of eukaryotic linear chromosomes, a process required for genomic stability and cell viability. Here we present the crystal structure of the active Tribolium castaneum telomerase catalytic subunit, TERT, bound to an RNA-DNA hairpin designed to resemble the putative RNA-templating region and telomeric DNA. The RNA-DNA hybrid adopts a helical structure, docked in the interior cavity of the TERT ring. Contacts between the RNA template and motifs 2 and B' position the solvent-accessible RNA bases close to the enzyme active site for nucleotide binding and selectivity. Nucleic acid binding induces rigid TERT conformational changes to form a tight catalytic complex. Overall, TERT-RNA template and TERT-telomeric DNA associations are remarkably similar to those observed for retroviral reverse transcriptases, suggesting common mechanistic aspects of DNA replication between the two families of enzymes. | ||
| - | |||
| - | Structural basis for telomerase catalytic subunit TERT binding to RNA template and telomeric DNA.,Mitchell M, Gillis A, Futahashi M, Fujiwara H, Skordalakes E Nat Struct Mol Biol. 2010 Apr;17(4):513-8. Epub 2010 Mar 28. PMID:20357774<ref>PMID:20357774</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 3kyl" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
| - | *[[Catalytic Subunit of T. Castaneum TERT Polymerase|Catalytic Subunit of T. Castaneum TERT Polymerase]] | ||
*[[Telomerase|Telomerase]] | *[[Telomerase|Telomerase]] | ||
*[[Telomerase 3D structures|Telomerase 3D structures]] | *[[Telomerase 3D structures|Telomerase 3D structures]] | ||
| - | == References == | ||
| - | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: | + | [[Category: Tribolium castaneum]] |
| - | [[Category: Skordalakes | + | [[Category: Skordalakes E]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
Structure of the catalytic subunit of telomerase bound to its RNA template and telomeric DNA
| |||||||||||