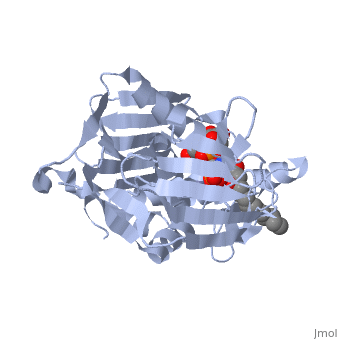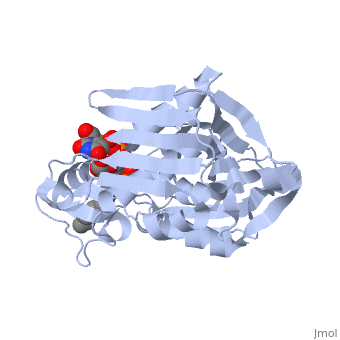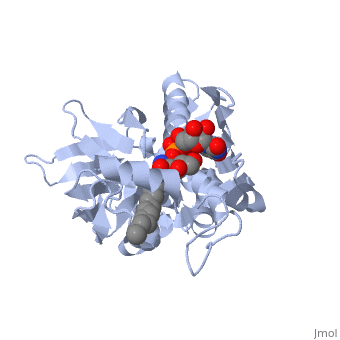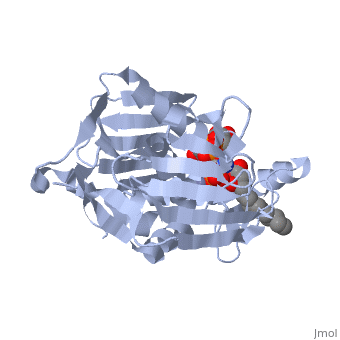
UDP-3-O-acyl-N-acetylglucosamine deacetylase
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 10: | Line 10: | ||
*<scene name='59/595824/Cv/6'>UDP-(3-O-(R-3-hydroxymyristoyl))-glucosamine binding site</scene>. | *<scene name='59/595824/Cv/6'>UDP-(3-O-(R-3-hydroxymyristoyl))-glucosamine binding site</scene>. | ||
*<scene name='59/595824/Cv/7'>Whole binding site</scene>. | *<scene name='59/595824/Cv/7'>Whole binding site</scene>. | ||
| + | *<scene name='59/595824/Cv/8'>Surface representation</scene>. | ||
| + | *<scene name='59/595824/Cv/9'>UDP-(3-O-(R-3-hydroxymyristoyl))-glucosamine is located in long tunnel</scene>. | ||
=== Para-(benzoyl)-phenylalanine as a potential inhibitor against LpxC of ''Leptospira spp.'': Homology modeling, docking and molecular dynamics study <ref>doi 10.1080/07391102.2012.758056</ref>=== | === Para-(benzoyl)-phenylalanine as a potential inhibitor against LpxC of ''Leptospira spp.'': Homology modeling, docking and molecular dynamics study <ref>doi 10.1080/07391102.2012.758056</ref>=== | ||
''Leptospira interrogans'', a Gram-negative bacterial pathogen is the main cause of human leptospirosis. Lipid A is a highly immunoreactive endotoxic center of lipopolysaccharide (LPS) that anchors LPS into the outer membrane of Leptospira. Discovery of compounds inhibiting lipid-A biosynthetic pathway would be promising for dissolving the structural integrity of membrane leading to cell lysis and death of Leptospira. <scene name='Journal:JBSD:10/Cv/2'>LpxC</scene>, a unique enzyme of lipid-A biosynthetic pathway was identified as common drug target of ''Leptospira''. Herein, homology modeling, docking, and molecular dynamics (MD) simulations were employed to discover potential inhibitors of LpxC. A <scene name='Journal:JBSD:10/Cv/3'>reliable tertiary structure of LpxC in complex with inhibitor BB-78485</scene> was constructed in Modeller 9v8. <scene name='Journal:JBSD:10/Cv/5'>Click here to see animated comparison</scene> between active site of target and template (LpxC from ''Pseudomonas aeruginosa'', PDB entry [[2ves]]) in the presence of BB-78485. The color code representations as <span style="color:deeppink;background-color:black;font-weight:bold;">deep pink: target</span>; <span style="color:lime;background-color:black;font-weight:bold;">green: template</span>; <span style="color:cyan;background-color:black;font-weight:bold;">cyan: BB-78485 in complex with target</span>; and <font color='darkmagenta'><b>darkmagenta: BB-78485 in complex with template</b></font>. Zn ion represented as sphere. A data-set of BB-78485 structural analogs were docked with LpxC in Maestro v9.2 virtual screening workflow, which implements three stage Glide docking protocol. Twelve lead molecules with better XP Gscore compared to BB-78485 were proposed as potential inhibitors of LpxC. Para-(benzoyl)-phenylalanine – that showed lowest XP Gscore (−10.35 kcal/mol) – was predicted to have best binding affinity towards LpxC. MD simulations were performed for LpxC and para-(benzoyl)-phenylalanine docking complex in Desmond v3.0. Trajectory analysis showed the docking complex and inter-molecular interactions was stable throughout the entire production part of MD simulations. The results indicate para-(benzoyl)-phenylalanine as a potent drug molecule against leptospirosis. | ''Leptospira interrogans'', a Gram-negative bacterial pathogen is the main cause of human leptospirosis. Lipid A is a highly immunoreactive endotoxic center of lipopolysaccharide (LPS) that anchors LPS into the outer membrane of Leptospira. Discovery of compounds inhibiting lipid-A biosynthetic pathway would be promising for dissolving the structural integrity of membrane leading to cell lysis and death of Leptospira. <scene name='Journal:JBSD:10/Cv/2'>LpxC</scene>, a unique enzyme of lipid-A biosynthetic pathway was identified as common drug target of ''Leptospira''. Herein, homology modeling, docking, and molecular dynamics (MD) simulations were employed to discover potential inhibitors of LpxC. A <scene name='Journal:JBSD:10/Cv/3'>reliable tertiary structure of LpxC in complex with inhibitor BB-78485</scene> was constructed in Modeller 9v8. <scene name='Journal:JBSD:10/Cv/5'>Click here to see animated comparison</scene> between active site of target and template (LpxC from ''Pseudomonas aeruginosa'', PDB entry [[2ves]]) in the presence of BB-78485. The color code representations as <span style="color:deeppink;background-color:black;font-weight:bold;">deep pink: target</span>; <span style="color:lime;background-color:black;font-weight:bold;">green: template</span>; <span style="color:cyan;background-color:black;font-weight:bold;">cyan: BB-78485 in complex with target</span>; and <font color='darkmagenta'><b>darkmagenta: BB-78485 in complex with template</b></font>. Zn ion represented as sphere. A data-set of BB-78485 structural analogs were docked with LpxC in Maestro v9.2 virtual screening workflow, which implements three stage Glide docking protocol. Twelve lead molecules with better XP Gscore compared to BB-78485 were proposed as potential inhibitors of LpxC. Para-(benzoyl)-phenylalanine – that showed lowest XP Gscore (−10.35 kcal/mol) – was predicted to have best binding affinity towards LpxC. MD simulations were performed for LpxC and para-(benzoyl)-phenylalanine docking complex in Desmond v3.0. Trajectory analysis showed the docking complex and inter-molecular interactions was stable throughout the entire production part of MD simulations. The results indicate para-(benzoyl)-phenylalanine as a potent drug molecule against leptospirosis. | ||
| - | </StructureSection> | ||
== 3D Structures of UDP-3-O-acyl-N-acetylglucosamine deacetylase == | == 3D Structures of UDP-3-O-acyl-N-acetylglucosamine deacetylase == | ||
| + | |||
| + | [[UDP-3-O-acyl-N-acetylglucosamine deacetylase 3D structures]] | ||
Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| Line 49: | Line 52: | ||
== References == | == References == | ||
<references/> | <references/> | ||
| - | + | </StructureSection> | |
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||