This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
3k9p
From Proteopedia
(Difference between revisions)
| (8 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | {{Seed}} | ||
| - | [[Image:3k9p.png|left|200px]] | ||
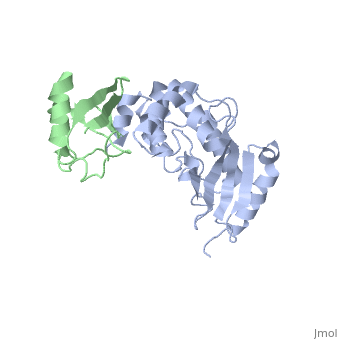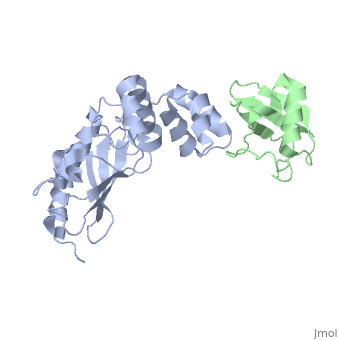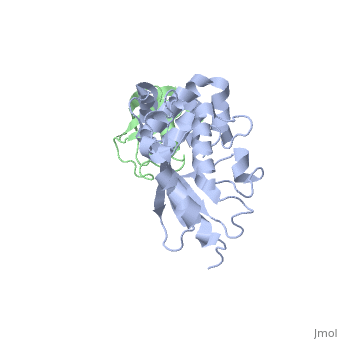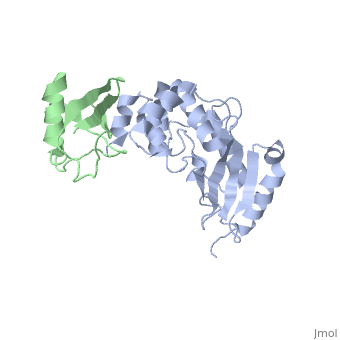
| - | + | ==The crystal structure of E2-25K and ubiquitin complex== | |
| - | + | <StructureSection load='3k9p' size='340' side='right'caption='[[3k9p]], [[Resolution|resolution]] 2.80Å' scene=''> | |
| - | + | == Structural highlights == | |
| - | or the | + | <table><tr><td colspan='2'>[[3k9p]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=3K9P OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=3K9P FirstGlance]. <br> |
| - | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.8Å</td></tr> | |
| - | --> | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=3k9p FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=3k9p OCA], [https://pdbe.org/3k9p PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=3k9p RCSB], [https://www.ebi.ac.uk/pdbsum/3k9p PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=3k9p ProSAT]</span></td></tr> |
| - | + | </table> | |
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/UBE2K_HUMAN UBE2K_HUMAN] Accepts ubiquitin from the E1 complex and catalyzes its covalent attachment to other proteins. In vitro, in the presence or in the absence of BRCA1-BARD1 E3 ubiquitin-protein ligase complex, catalyzes the synthesis of 'Lys-48'-linked polyubiquitin chains. Does not transfer ubiquitin directly to but elongates monoubiquitinated substrate protein. Mediates the selective degradation of short-lived and abnormal proteins, such as the endoplasmic reticulum-associated degradation (ERAD) of misfolded lumenal proteins. Ubiquitinates huntingtin. May mediate foam cell formation by the suppression of apoptosis of lipid-bearing macrophages through ubiquitination and subsequence degradation of p53/TP53. Proposed to be involved in ubiquitination and proteolytic processing of NF-kappa-B; in vitro supports ubiquitination of NFKB1. In case of infection by cytomegaloviruses may be involved in the US11-dependent degradation of MHC class I heavy chains following their export from the ER to the cytosol. In case of viral infections may be involved in the HPV E7 protein-dependent degradation of RB1.<ref>PMID:8702625</ref> <ref>PMID:10634809</ref> <ref>PMID:10675012</ref> <ref>PMID:16714285</ref> <ref>PMID:16868077</ref> <ref>PMID:17873885</ref> <ref>PMID:20061386</ref> <ref>PMID:19906396</ref> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/k9/3k9p_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=3k9p ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| - | == | + | ==See Also== |
| - | + | *[[Ubiquitin|Ubiquitin]] | |
| - | + | *[[3D structures of ubiquitin|3D structures of ubiquitin]] | |
| - | + | *[[3D structures of ubiquitin conjugating enzyme|3D structures of ubiquitin conjugating enzyme]] | |
| - | + | == References == | |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | == | + | |
| - | < | + | |
[[Category: Homo sapiens]] | [[Category: Homo sapiens]] | ||
| - | [[Category: | + | [[Category: Large Structures]] |
| - | [[Category: Eom | + | [[Category: Eom SH]] |
| - | [[Category: Kang | + | [[Category: Kang GB]] |
| - | [[Category: Ko | + | [[Category: Ko S]] |
| - | [[Category: Lee | + | [[Category: Lee W]] |
| - | [[Category: Song | + | [[Category: Song SM]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
The crystal structure of E2-25K and ubiquitin complex
| |||||||||||
Categories: Homo sapiens | Large Structures | Eom SH | Kang GB | Ko S | Lee W | Song SM