This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1kcg
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='1kcg' size='340' side='right'caption='[[1kcg]], [[Resolution|resolution]] 2.60Å' scene=''> | <StructureSection load='1kcg' size='340' side='right'caption='[[1kcg]], [[Resolution|resolution]] 2.60Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1kcg]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/ | + | <table><tr><td colspan='2'>[[1kcg]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1KCG OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1KCG FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=GSH:GLUTATHIONE'>GSH</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.6Å</td></tr> |
| + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=GSH:GLUTATHIONE'>GSH</scene></td></tr> | ||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1kcg FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1kcg OCA], [https://pdbe.org/1kcg PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1kcg RCSB], [https://www.ebi.ac.uk/pdbsum/1kcg PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1kcg ProSAT]</span></td></tr> | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1kcg FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1kcg OCA], [https://pdbe.org/1kcg PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1kcg RCSB], [https://www.ebi.ac.uk/pdbsum/1kcg PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1kcg ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
| - | + | [https://www.uniprot.org/uniprot/NKG2D_HUMAN NKG2D_HUMAN] Receptor for MICA, MICB, ULBP1, ULBP2, ULBP3 (ULBP2>ULBP1>ULBP3) and ULBP4. Plays a role as a receptor for the recognition of MHC class I HLA-E molecules by NK cells and some cytotoxic T-cells. Involved in the immune surveillance exerted by T- and B-lymphocytes. | |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 19: | Line 20: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1kcg ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1kcg ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
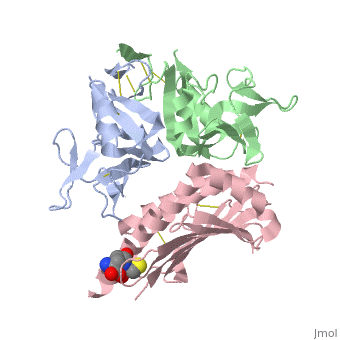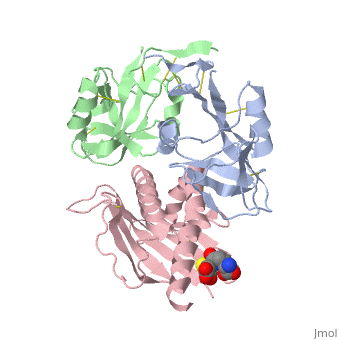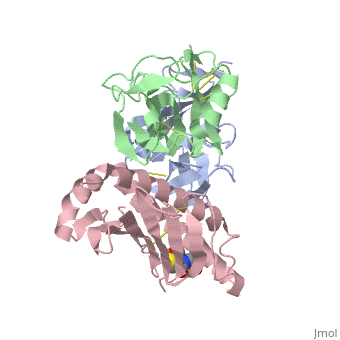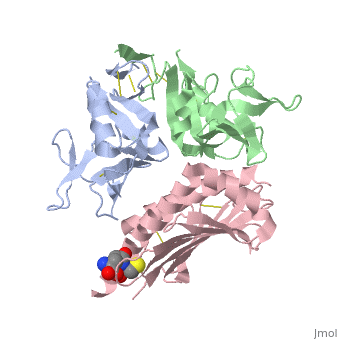
| - | NKG2D is known to trigger the natural killer (NK) cell lysis of various tumor and virally infected cells. In the NKG2D/ULBP3 complex, the structure of ULBP3 resembles the alpha1 and alpha2 domains of classical MHC molecules without a bound peptide. The lack of alpha3 and beta2m domains is compensated by replacing two hydrophobic patches at the underside of the class I MHC-like beta sheet floor with a group of hydrophilic and charged residues in ULBP3. NKG2D binds diagonally across the ULBP3 alpha helices, creating a complementary interface, an asymmetrical subunit orientation, and local conformational adjustments in the receptor. The interface is stabilized primarily by hydrogen bonds and hydrophobic interactions. Unlike the KIR receptors that recognize a conserved HLA region by a lock-and-key mechanism, NKG2D recognizes diverse ligands by an induced-fit mechanism. | ||
| - | |||
| - | Conformational plasticity revealed by the cocrystal structure of NKG2D and its class I MHC-like ligand ULBP3.,Radaev S, Rostro B, Brooks AG, Colonna M, Sun PD Immunity. 2001 Dec;15(6):1039-49. PMID:11754823<ref>PMID:11754823</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 1kcg" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
*[[NK cell receptor|NK cell receptor]] | *[[NK cell receptor|NK cell receptor]] | ||
| - | == References == | ||
| - | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
| - | [[Category: | + | [[Category: Homo sapiens]] |
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: Radaev | + | [[Category: Radaev S]] |
| - | [[Category: Sun | + | [[Category: Sun P]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
NKG2D in complex with ULBP3
| |||||||||||