Glutamate dehydrogenase
From Proteopedia
(Difference between revisions)
| (25 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
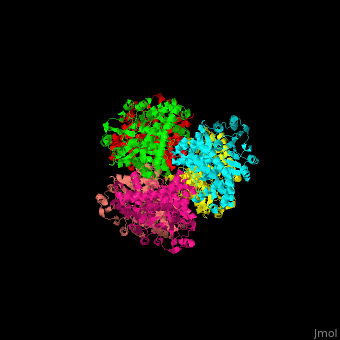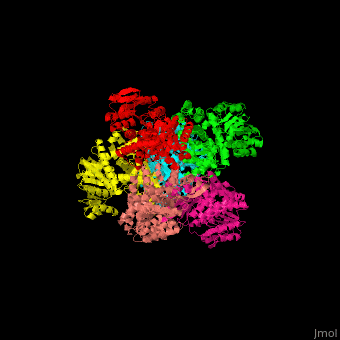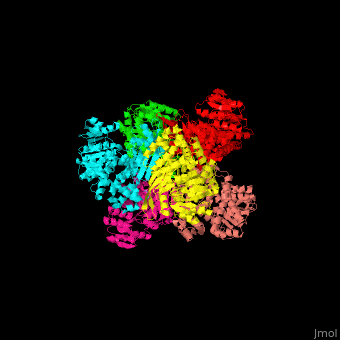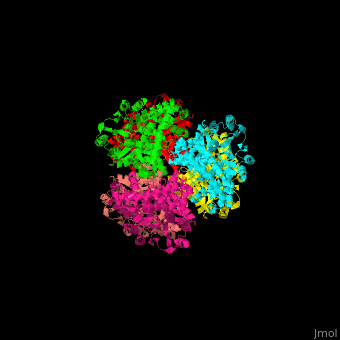
| - | + | <StructureSection load='' size='350' side='right' caption='Glutamate dehydrogenase complex with glutamate (PDB entry [[1bgv]])' scene='49/490918/Cv/3'> | |
| - | + | == Function == | |
| - | + | '''Glutamate dehydrogenase''' (GLDH) catalyzes the reversible conversion of glutamate to α-ketoglutarate (AKG) and ammonium. The ammonia is removed via the urea cycle. [[NAD]] or NADP is a cofactor in GLDH activity. NAD is a cofactor in the forward reaction while NADP is a cofactor in the reverse reaction. GLDH is regulated by the cell’s energy state. ATP and GTP inhibit the enzyme while ADP, GDP and leucine positively enhance it<ref>PMID:14299621</ref>. '''Glutamate dehydrogenase 1''' (GLDH1) catalyzes the deamination of glutamate to 2-oxoglutarate and ammonium. GLDH1 is regulated in the same manner as GLDH<ref>PMID:12054821</ref>. | |
| - | + | See also [[Citric Acid Cycle]]. | |
| - | + | ||
| - | + | ||
| - | '''Glutamate dehydrogenase''' (GLDH) catalyzes the reversible conversion of glutamate to α-ketoglutarate and ammonium. The ammonia is removed via the urea cycle. | + | |
| - | == | + | == Relevance == |
| + | Elevated GLDH values in blood serum indicate liver malfunction<ref>PMID:589307</ref>. | ||
| - | + | == Disease == | |
| + | Mutations in GLDH1 are associated with familial hyperinsulinism<ref>PMID:10453735</ref>. | ||
| - | + | == Structural highlights == | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | ''' | + | The biological assembly of GLDH from ''Clostridium symbiosum'' is <scene name='49/490918/Cv/6'>homohexamer</scene>. The <scene name='49/490918/Cv/7'>glutamate binding pocket of GLDH</scene> is in a cleft between the two domains of the enzyme with <scene name='49/490918/Cv/8'>residue D165 serving as a general base and a protein-bound water molecule as the attacking nucleophile</scene> <ref>PMID:8263917</ref>. Water molecules are shown as red spheres. |
| + | |||
| + | ==3D structures of glutamate dehydrogenase== | ||
| + | [[Glutamate dehydrogenase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | == References == | ||
| + | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ FRIEDEN C. GLUTAMATE DEHYDROGENASE. VI. SURVEY OF PURINE NUCLEOTIDE AND OTHER EFFECTS ON THE ENZYME FROM VARIOUS SOURCES. J Biol Chem. 1965 May;240:2028-35. PMID:14299621
- ↑ Smith TJ, Schmidt T, Fang J, Wu J, Siuzdak G, Stanley CA. The structure of apo human glutamate dehydrogenase details subunit communication and allostery. J Mol Biol. 2002 May 3;318(3):765-77. PMID:12054821 doi:10.1016/S0022-2836(02)00161-4
- ↑ Van Waes L, Lieber CS. Glutamate dehydrogenase: a reliable marker of liver cell necrosis in the alcoholic. Br Med J. 1977 Dec 10;2(6101):1508-10. PMID:589307
- ↑ Yorifuji T, Muroi J, Uematsu A, Hiramatsu H, Momoi T. Hyperinsulinism-hyperammonemia syndrome caused by mutant glutamate dehydrogenase accompanied by novel enzyme kinetics. Hum Genet. 1999 Jun;104(6):476-9. PMID:10453735
- ↑ Stillman TJ, Baker PJ, Britton KL, Rice DW. Conformational flexibility in glutamate dehydrogenase. Role of water in substrate recognition and catalysis. J Mol Biol. 1993 Dec 20;234(4):1131-9. PMID:8263917 doi:http://dx.doi.org/10.1006/jmbi.1993.1665