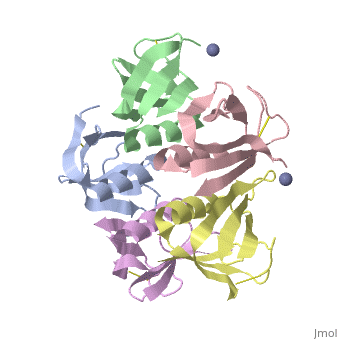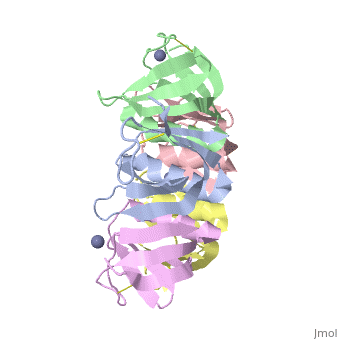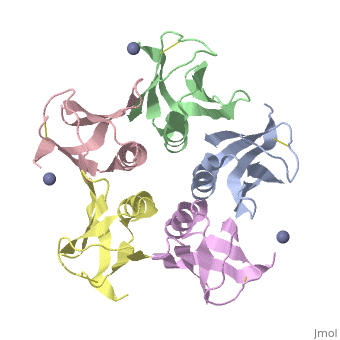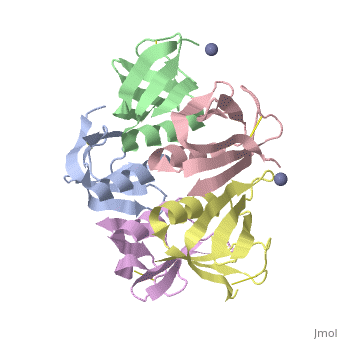This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2xsc
From Proteopedia
(Difference between revisions)
| (9 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | {{Seed}} | ||
| - | [[Image:2xsc.jpg|left|200px]] | ||
| - | + | ==Crystal structure of the cell-binding B oligomer of verotoxin-1 from E. coli== | |
| - | + | <StructureSection load='2xsc' size='340' side='right'caption='[[2xsc]], [[Resolution|resolution]] 2.05Å' scene=''> | |
| - | + | == Structural highlights == | |
| - | + | <table><tr><td colspan='2'>[[2xsc]] is a 5 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_coli Escherichia coli]. This structure supersedes the now removed PDB entry [http://oca.weizmann.ac.il/oca-bin/send-pdb?obs=1&id=1bov 1bov]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2XSC OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2XSC FirstGlance]. <br> | |
| - | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.052Å</td></tr> | |
| - | -- | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=ZN:ZINC+ION'>ZN</scene></td></tr> |
| - | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2xsc FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2xsc OCA], [https://pdbe.org/2xsc PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2xsc RCSB], [https://www.ebi.ac.uk/pdbsum/2xsc PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2xsc ProSAT]</span></td></tr> | |
| + | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/STXB_BPH19 STXB_BPH19] The B subunit is responsible for the binding of the holotoxin to specific receptors on the target cell surface, such as globotriaosylceramide (Gb3) in human intestinal microvilli. | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | The Shiga toxin family, a group of cytotoxins associated with diarrhoeal diseases and the haemolytic uraemic syndrome, includes Shiga toxin from Shigella dysenteriae type 1 and verotoxins produced by enteropathogenic Escherichia coli. The family belongs to the A-B class of bacterial toxins, which includes the cholera toxin family, pertussis and diphtheria toxins. These toxins all have bipartite structures consisting of an enzymatic A subunit associated with a B oligomer which binds to specific cell-surface receptors, but their amino-acid sequences and pathogenic mechanisms differ. We have determined the crystal structure of the B oligomer of verotoxin-1 from E. coli. The structure unexpectedly resembles that of the B oligomer of the cholera toxin-like heat-labile enterotoxin from E. coli, despite the absence of detectable sequence similarity between these two proteins. This result implies a distant evolutionary relationship between the Shiga toxin and cholera toxin families. We suggest that the cell surface receptor-binding site lies in a cleft between adjacent subunits of the B pentamer, providing a potential target for drugs and vaccines to prevent toxin binding and effect. | ||
| - | + | Crystal structure of the cell-binding B oligomer of verotoxin-1 from E. coli.,Stein PE, Boodhoo A, Tyrrell GJ, Brunton JL, Read RJ Nature. 1992 Feb 20;355(6362):748-50. PMID:1741063<ref>PMID:1741063</ref> | |
| + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| + | </div> | ||
| + | <div class="pdbe-citations 2xsc" style="background-color:#fffaf0;"></div> | ||
| - | + | ==See Also== | |
| - | + | *[[Shiga toxin|Shiga toxin]] | |
| - | + | *[[Shiga toxin 3D structures|Shiga toxin 3D structures]] | |
| - | + | == References == | |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | == | + | </StructureSection> |
| - | + | ||
| - | + | ||
| - | == | + | |
| - | < | + | |
[[Category: Escherichia coli]] | [[Category: Escherichia coli]] | ||
| - | [[Category: Boodhoo | + | [[Category: Large Structures]] |
| - | [[Category: Brunton | + | [[Category: Boodhoo A]] |
| - | [[Category: Bunkoczi | + | [[Category: Brunton JL]] |
| - | [[Category: Oeffner | + | [[Category: Bunkoczi G]] |
| - | [[Category: Read | + | [[Category: Oeffner RD]] |
| - | [[Category: Stein | + | [[Category: Read RJ]] |
| - | [[Category: Tyrrell | + | [[Category: Stein PE]] |
| - | + | [[Category: Tyrrell GJ]] | |
| - | + | ||
| - | + | ||
Current revision
Crystal structure of the cell-binding B oligomer of verotoxin-1 from E. coli
| |||||||||||