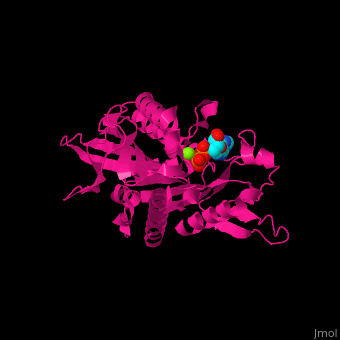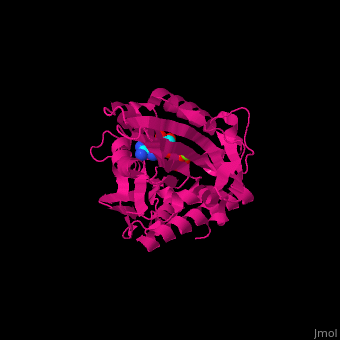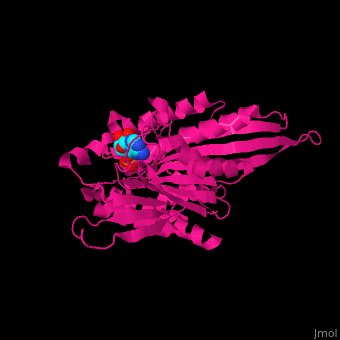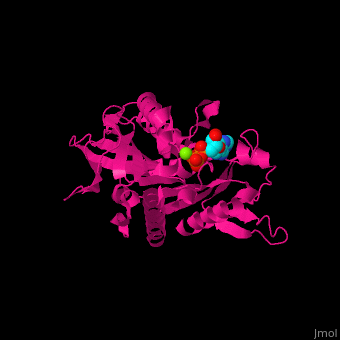This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Kinesin
From Proteopedia
(Difference between revisions)
| (51 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | [[ | + | <StructureSection load='' size='350' side='right' scene='41/410296/Cv/1' caption='KIF1A Motor Domain complex with ADP and Mg+2 ion (green), [[2zfi]]'> |
| - | + | == Function == | |
| - | ''' | + | [[Kinesin|Kinesins]] are eukaryotic motor proteins which move along microtubules<ref>PMID:19773780</ref>. Kinesin (KIF) is a dimer consisting of 2 heavy chains and two light chains. The heavy chain contains the N-terminal globular motor domain (MD) residues 1-375, responsible for the motor activity of kinesin, a central flexible neck linker (FNL) coiled-coil stalk which intertwines to form the dimer and a small globular C-terminal domain which interacts with other proteins like the kinesin light chain. The light chain (KLC) forms the tail region. The KLC contains a cargo binding domain which is called TPR (Tetratricopeptide repeat). The KIFs are named by their gene number. KIF are organized into 14 families named kinesin-1 to kinesin-14. KIF contains a forkhead-associated domain (FHA) which is involved in phosphopeptide recognition.<br /> |
| - | + | *'''KIF1A''' transports organelles along axonal microtubules.<br /> | |
| + | *'''KIF1C''' and '''KIF2''' are plus-ended directed microtubule motors.<br /> | ||
| + | *'''KIF2C''', '''KIF22''' and '''KIF3B''' are plus-ended directed microtubule motors in mitotic cells<br /> | ||
| + | *'''KIF13B''' transports VEGFR2 from the Golgi to the endothelial cell surface.<br /> | ||
| + | *'''KIF16B''' is plus-ended directed microtubule motor involved in endosome transport and receptor recycling.<br /> | ||
| + | *'''KIFC1''' transports DNA molecules along cytoskeleton filaments.<br /> | ||
| + | *'''Kar3''' is kinesin protein in yeast.<br /> | ||
| + | *'''Eg5''' or '''KIF11''' is a kinesin (See [[Kinesin-5]]) which participates in mitosis.<br /> | ||
| + | *'''NOD''' is a ''Drosophila'' chromosome-associated kinesin. | ||
| + | See also [[CAP-Gly domain]]. | ||
| - | == | + | == Disease == |
| - | + | Mutations in KIF5A are involved in hereditary spastic paraplegia<ref>PMID:18203753</ref>. Mutation in KIF1B is the cause of Charcot-Marie-Tooth disease <ref>PMID:14559185</ref>. Mutations in KIF22 cause spondyloepimetaphyseal dysplasia<ref>PMID:23665482</ref>. | |
| - | + | ||
| - | + | ||
| + | == Structural highlights == | ||
| + | <scene name='41/410296/Cv/10'>Active site</scene>. Water molecules shown as red spheres. Residue <scene name='41/410296/Cv/11'>Arg216 is the key residue in KIF for the chemical cycling of ATPase and for the mechanical cycling</scene>. Arg216 pivots to enable <scene name='41/410296/Cv/12'>Mg-ADP</scene> release or the phosphate release. <scene name='41/410296/Cv/13'>Arg216 forms a latch</scene> in the KIF 'closed-state' before the Mg-ADP release. Binding of β-tubulin to KIF releases the latch, enabling the KIF conformation change and detaching KIF from the microtubule and enabling the next movement cycle<ref>PMID:18806800</ref>. | ||
| + | <scene name='41/410296/Cv/14'>Mg coordination site</scene>. | ||
== 3D Structures of Kinesin == | == 3D Structures of Kinesin == | ||
| + | [[Kinesin 3D Structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | == References == | |
| - | + | <references/> | |
| - | [[ | + | [[Category:Topic Page]] |
Current revision
| |||||||||||
References
- ↑ Hirokawa N, Noda Y, Tanaka Y, Niwa S. Kinesin superfamily motor proteins and intracellular transport. Nat Rev Mol Cell Biol. 2009 Oct;10(10):682-96. doi: 10.1038/nrm2774. PMID:19773780 doi:http://dx.doi.org/10.1038/nrm2774
- ↑ Ebbing B, Mann K, Starosta A, Jaud J, Schols L, Schule R, Woehlke G. Effect of spastic paraplegia mutations in KIF5A kinesin on transport activity. Hum Mol Genet. 2008 May 1;17(9):1245-52. doi: 10.1093/hmg/ddn014. Epub 2008 Jan, 18. PMID:18203753 doi:http://dx.doi.org/10.1093/hmg/ddn014
- ↑ Hirokawa N, Takemura R. Biochemical and molecular characterization of diseases linked to motor proteins. Trends Biochem Sci. 2003 Oct;28(10):558-65. PMID:14559185 doi:http://dx.doi.org/10.1016/j.tibs.2003.08.006
- ↑ Grosch M, Gruner B, Spranger S, Stutz AM, Rausch T, Korbel JO, Seelow D, Nurnberg P, Sticht H, Lausch E, Zabel B, Winterpacht A, Tagariello A. Identification of a Ninein (NIN) mutation in a family with spondyloepimetaphyseal dysplasia with joint laxity (leptodactylic type)-like phenotype. Matrix Biol. 2013 Oct-Nov;32(7-8):387-92. doi: 10.1016/j.matbio.2013.05.001. Epub, 2013 May 9. PMID:23665482 doi:http://dx.doi.org/10.1016/j.matbio.2013.05.001
- ↑ Nitta R, Okada Y, Hirokawa N. Structural model for strain-dependent microtubule activation of Mg-ADP release from kinesin. Nat Struct Mol Biol. 2008 Oct;15(10):1067-75. Epub 2008 Sep 21. PMID:18806800 doi:10.1038/nsmb.1487
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, Jaime Prilusky