This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
TonB
From Proteopedia
| (12 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
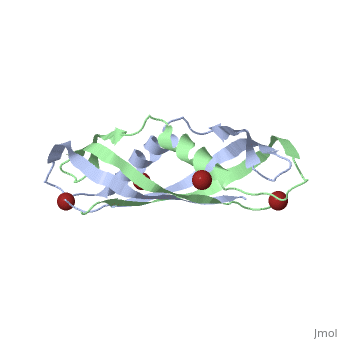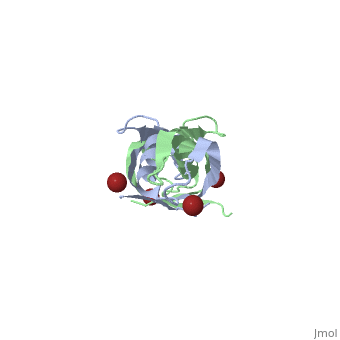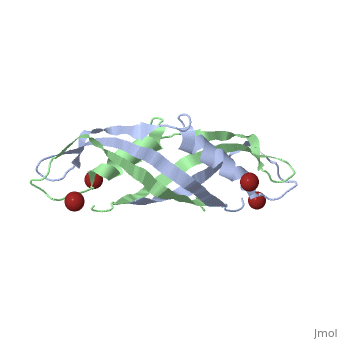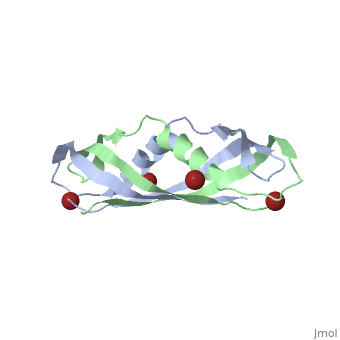
| - | + | <StructureSection load='1ihr' size='350' side='right' scene='' caption='E. coli TonB C-terminal dimer complex with Br- ions (dark red) (PDB code [[1ihr]])'> | |
| - | + | ||
==Structure== | ==Structure== | ||
| - | + | '''TonB''' spans the periplasm <ref>PMID: 16741125</ref>, with the C-terminus of TonB spanning from residues ~150 to 239<ref name='Postle'>PMID: 21179522</ref>. For more details on the Ton system see [[Ton]]. | |
==Function== | ==Function== | ||
| Line 17: | Line 16: | ||
===BtuB-TonB Complex=== | ===BtuB-TonB Complex=== | ||
| - | + | <StructureSection load='2gsk' size='350' side='right' scene='BtuB-TonB_Complex/Btubtonbcomplex/1' caption='E. coli TonB C-terminal (green) complex with vitamin B12 transporter BtuB (grey) (PDB code [[2gsk]])'> | |
| - | + | ||
As shown in the 3D structure to the right (2GSK), TonB complexes with [[BtuB]] in order to aid the transport of nutrients such as cobalamins<ref name='Cadieux'>PMID 11029413</ref> across the outer membrane by incorporating the proton-motive force into the outer membrane.<ref name='Shultis'>PMID: 16741124</ref> TonB attaches to BtuB on the periplasmic side of the 614 amino acid BtuB protein<ref name='Shultis'> PMID: 16741124</ref>. | As shown in the 3D structure to the right (2GSK), TonB complexes with [[BtuB]] in order to aid the transport of nutrients such as cobalamins<ref name='Cadieux'>PMID 11029413</ref> across the outer membrane by incorporating the proton-motive force into the outer membrane.<ref name='Shultis'>PMID: 16741124</ref> TonB attaches to BtuB on the periplasmic side of the 614 amino acid BtuB protein<ref name='Shultis'> PMID: 16741124</ref>. | ||
| + | </StructureSection> | ||
| + | ==3D structures of TonB== | ||
| + | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| + | |||
| + | [[1ihr]], [[1qxx]], [[1u07]] – EcTonB C terminal – ''Escherichia coli''<br /> | ||
| + | [[1xx3]], [[5lw8]] - EcTonB C terminal – NMR<br /> | ||
| + | [[2grx]] - EcTonB + ferrichrome-iron receptor<br /> | ||
| + | [[2ivz]] - EcTonB + colicin-E9 peptide<br /> | ||
| + | [[2gsk]] - EcTonB C terminal + vitamin B12 transporter BtuB<br /> | ||
| + | [[2k9k]] – TonB2 C terminal – ''Listonella anguillarum''<br /> | ||
| + | [[6fip]] - PaTonB C terminal – ''Pseudomonas aeruginosa'' - NMR<br /> | ||
| + | [[6i97]] - PaTonB C terminal + TonB-dependent receptor <br /> | ||
| + | [[5lw8]], [[6sly]] - TonB C terminal – ''Helicobacter pylori'' - NMR<br /> | ||
== References== | == References== | ||
<references/> | <references/> | ||
| + | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D structures of TonB
Updated on 06-December-2020
1ihr, 1qxx, 1u07 – EcTonB C terminal – Escherichia coli
1xx3, 5lw8 - EcTonB C terminal – NMR
2grx - EcTonB + ferrichrome-iron receptor
2ivz - EcTonB + colicin-E9 peptide
2gsk - EcTonB C terminal + vitamin B12 transporter BtuB
2k9k – TonB2 C terminal – Listonella anguillarum
6fip - PaTonB C terminal – Pseudomonas aeruginosa - NMR
6i97 - PaTonB C terminal + TonB-dependent receptor
5lw8, 6sly - TonB C terminal – Helicobacter pylori - NMR
References
- ↑ Pawelek PD, Croteau N, Ng-Thow-Hing C, Khursigara CM, Moiseeva N, Allaire M, Coulton JW. Structure of TonB in complex with FhuA, E. coli outer membrane receptor. Science. 2006 Jun 2;312(5778):1399-402. PMID:16741125 doi:312/5778/1399
- ↑ 2.0 2.1 Postle K, Kastead KA, Gresock MG, Ghosh J, Swayne CD. The TonB Dimeric Crystal Structures Do Not Exist In Vivo. MBio. 2010 Dec 21;1(5). pii: e00307-10. PMID:21179522 doi:10.1128/mBio.00307-10
- ↑ 3.0 3.1 Kampfenkel K, Braun V. Topology of the ExbB protein in the cytoplasmic membrane of Escherichia coli. J Biol Chem. 1993 Mar 15;268(8):6050-7. PMID:8449962
- ↑ Wiener MC. TonB-dependent outer membrane transport: going for Baroque? Curr Opin Struct Biol. 2005 Aug;15(4):394-400. PMID:16039843 doi:10.1016/j.sbi.2005.07.001
- ↑ Held KG, Postle K. ExbB and ExbD do not function independently in TonB-dependent energy transduction. J Bacteriol. 2002 Sep;184(18):5170-3. PMID:12193634
- ↑ Cadieux N, Bradbeer C, Kadner RJ. Sequence changes in the ton box region of BtuB affect its transport activities and interaction with TonB protein. J Bacteriol. 2000 Nov;182(21):5954-61. PMID:11029413
- ↑ 7.0 7.1 Shultis DD, Purdy MD, Banchs CN, Wiener MC. Outer membrane active transport: structure of the BtuB:TonB complex. Science. 2006 Jun 2;312(5778):1396-9. PMID:16741124 doi:312/5778/1396