This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Crystal structure of protein from Staphylococcus aureus
From Proteopedia
| (One intermediate revision not shown.) | |||
| Line 1: | Line 1: | ||
| + | <StructureSection load='1tsj' size='400' side='right' scene= caption=''> | ||
| + | |||
[[Image:1tsj.png|left|200px]] | [[Image:1tsj.png|left|200px]] | ||
| - | + | =Crystal structure of hypothetical protein from Staphylococcus aureus ([[1tsj]])= | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | =Crystal structure of protein from Staphylococcus aureus= | + | |
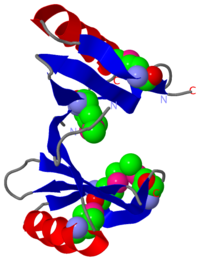
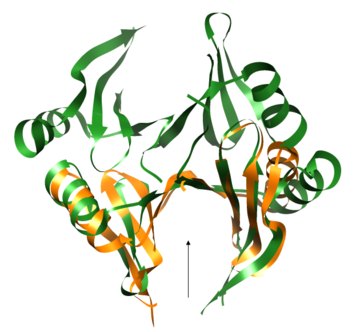
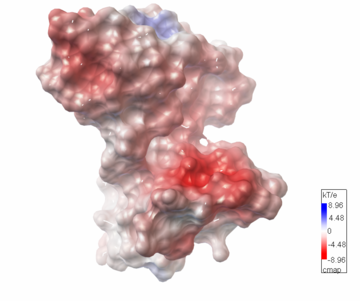
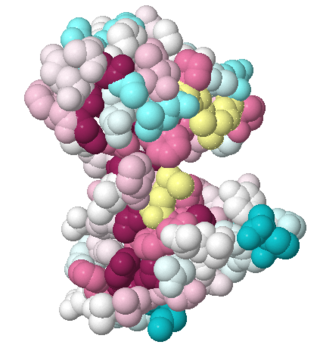
| - | PDB entry 1tsj refers to a hypothetical protein of 139 residues which is predicted as both a dimer<ref> PMID:9787643</ref> and a cytoplasmic protein.<ref>PMID:15699023</ref><ref>PMID:12824378</ref><ref>PMID:14990451</ref><ref>PMID:11524373</ref> | + | PDB entry [[1tsj]] refers to a hypothetical protein of 139 residues which is predicted as both a dimer<ref> PMID:9787643</ref> and a cytoplasmic protein.<ref>PMID:15699023</ref><ref>PMID:12824378</ref><ref>PMID:14990451</ref><ref>PMID:11524373</ref> |
The protein is associated with Pfam<ref>PMID:18428696</ref> entry PF06983 of 3-demethylubiquinone-9 3-methyltransferases. | The protein is associated with Pfam<ref>PMID:18428696</ref> entry PF06983 of 3-demethylubiquinone-9 3-methyltransferases. | ||
| Line 48: | Line 32: | ||
It seems more likely that 1tsj belongs to the Glyoxalase/bleomycin resistance protein/dioxygenase superfamily and functions as a demethylubiquinone-9 3-methyltransferase protein than that it functions as Oxidoreductase. | It seems more likely that 1tsj belongs to the Glyoxalase/bleomycin resistance protein/dioxygenase superfamily and functions as a demethylubiquinone-9 3-methyltransferase protein than that it functions as Oxidoreductase. | ||
| + | </StructureSection> | ||
Current revision
| |||||||||||
References
- ↑ Henrick K, Thornton JM. PQS: a protein quaternary structure file server. Trends Biochem Sci. 1998 Sep;23(9):358-61. PMID:9787643
- ↑ Bhasin M, Garg A, Raghava GP. PSLpred: prediction of subcellular localization of bacterial proteins. Bioinformatics. 2005 May 15;21(10):2522-4. Epub 2005 Feb 4. PMID:15699023 doi:10.1093/bioinformatics/bti309
- ↑ Gardy JL, Spencer C, Wang K, Ester M, Tusnady GE, Simon I, Hua S, deFays K, Lambert C, Nakai K, Brinkman FS. PSORT-B: Improving protein subcellular localization prediction for Gram-negative bacteria. Nucleic Acids Res. 2003 Jul 1;31(13):3613-7. PMID:12824378
- ↑ Lu Z, Szafron D, Greiner R, Lu P, Wishart DS, Poulin B, Anvik J, Macdonell C, Eisner R. Predicting subcellular localization of proteins using machine-learned classifiers. Bioinformatics. 2004 Mar 1;20(4):547-56. Epub 2004 Jan 22. PMID:14990451 doi:10.1093/bioinformatics/bth026
- ↑ Hua S, Sun Z. Support vector machine approach for protein subcellular localization prediction. Bioinformatics. 2001 Aug;17(8):721-8. PMID:11524373
- ↑ Finn R, Griffiths-Jones S, Bateman A. Identifying protein domains with the Pfam database. Curr Protoc Bioinformatics. 2003 May;Chapter 2:Unit 2.5. PMID:18428696 doi:10.1002/0471250953.bi0205s01
- ↑ Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997 Sep 1;25(17):3389-402. PMID:9254694
- ↑ Arnold K, Kiefer F, Kopp J, Battey JN, Podvinec M, Westbrook JD, Berman HM, Bordoli L, Schwede T. The protein model portal. J Struct Funct Genomics. 2009 Mar;10(1):1-8. Epub 2008 Nov 27. PMID:19037750 doi:10.1007/s10969-008-9048-5
- ↑ Emmert DB, Stoehr PJ, Stoesser G, Cameron GN. The European Bioinformatics Institute (EBI) databases. Nucleic Acids Res. 1994 Sep;22(17):3445-9. PMID:7937043
- ↑ Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem. 2004 Oct;25(13):1605-12. PMID:15264254 doi:10.1002/jcc.20084
- ↑ Liang J, Edelsbrunner H, Woodward C. Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Protein Sci. 1998 Sep;7(9):1884-97. PMID:9761470
- ↑ Thornalley PJ. Glyoxalase I--structure, function and a critical role in the enzymatic defence against glycation. Biochem Soc Trans. 2003 Dec;31(Pt 6):1343-8. PMID:14641060 doi:10.1042/
- ↑ Sanner MF. Python: a programming language for software integration and development. J Mol Graph Model. 1999 Feb;17(1):57-61. PMID:10660911
- ↑ Goldenberg O, Erez E, Nimrod G, Ben-Tal N. The ConSurf-DB: pre-calculated evolutionary conservation profiles of protein structures. Nucleic Acids Res. 2009 Jan;37(Database issue):D323-7. Epub 2008 Oct 29. PMID:18971256 doi:http://dx.doi.org/10.1093/nar/gkn822
Created with the participation of Talya Etzion.
Categories: Staphylococcus aureus subsp. aureus | Burley, S K. | Gorman, J. | Min, T. | NYSGXRC, New York Structural GenomiX Research Consortium. | Shapiro, L. | Conserved hypothetical protein | Crystal structure | New york structural genomics consortium | New york structural genomix research consortium | Nysgxrc | Protein structure initiative | Psi | Structural genomic