1z1g
From Proteopedia
(Difference between revisions)
| (9 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | [[Image:1z1g.png|left|200px]] | ||
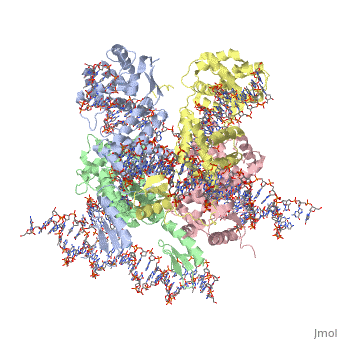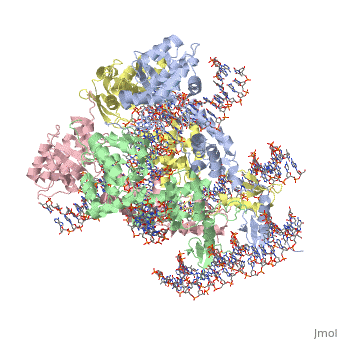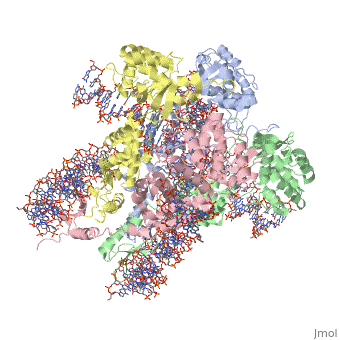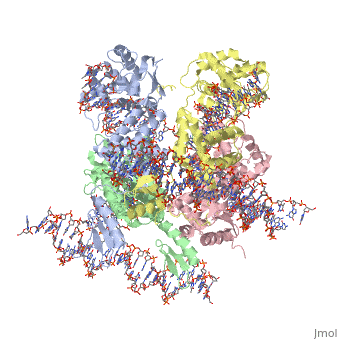
| - | + | ==Crystal structure of a lambda integrase tetramer bound to a Holliday junction== | |
| + | <StructureSection load='1z1g' size='340' side='right'caption='[[1z1g]], [[Resolution|resolution]] 4.40Å' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[1z1g]] is a 12 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_virus_Lambda Escherichia virus Lambda]. The December 2006 RCSB PDB [https://pdb.rcsb.org/pdb/static.do?p=education_discussion/molecule_of_the_month/index.html Molecule of the Month] feature on ''Transposase'' by David S. Goodsell is [https://dx.doi.org/10.2210/rcsb_pdb/mom_2006_12 10.2210/rcsb_pdb/mom_2006_12]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1Z1G OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1Z1G FirstGlance]. <br> | ||
| + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 4.4Å</td></tr> | ||
| + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=MSE:SELENOMETHIONINE'>MSE</scene></td></tr> | ||
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1z1g FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1z1g OCA], [https://pdbe.org/1z1g PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1z1g RCSB], [https://www.ebi.ac.uk/pdbsum/1z1g PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1z1g ProSAT]</span></td></tr> | ||
| + | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/VINT_LAMBD VINT_LAMBD] Integrase is necessary for integration of the phage into the host genome by site-specific recombination. In conjunction with excisionase, integrase is also necessary for excision of the prophage from the host genome. | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/z1/1z1g_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1z1g ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | Site-specific DNA recombination is important for basic cellular functions including viral integration, control of gene expression, production of genetic diversity and segregation of newly replicated chromosomes, and is used by bacteriophage lambda to integrate or excise its genome into and out of the host chromosome. lambda recombination is carried out by the bacteriophage-encoded integrase protein (lambda-int) together with accessory DNA sites and associated bending proteins that allow regulation in response to cell physiology. Here we report the crystal structures of lambda-int in higher-order complexes with substrates and regulatory DNAs representing different intermediates along the reaction pathway. The structures show how the simultaneous binding of two separate domains of lambda-int to DNA facilitates synapsis and can specify the order of DNA strand cleavage and exchange. An intertwined layer of amino-terminal domains bound to accessory (arm) DNAs shapes the recombination complex in a way that suggests how arm binding shifts the reaction equilibrium in favour of recombinant products. | ||
| - | + | A structural basis for allosteric control of DNA recombination by lambda integrase.,Biswas T, Aihara H, Radman-Livaja M, Filman D, Landy A, Ellenberger T Nature. 2005 Jun 23;435(7045):1059-66. PMID:15973401<ref>PMID:15973401</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | <div class="pdbe-citations 1z1g" style="background-color:#fffaf0;"></div> | |
| - | + | ||
==See Also== | ==See Also== | ||
*[[Phage integrase|Phage integrase]] | *[[Phage integrase|Phage integrase]] | ||
| - | *[[ | + | *[[Retroviral integrase 3D structures|Retroviral integrase 3D structures]] |
| - | + | == References == | |
| - | == | + | <references/> |
| - | < | + | __TOC__ |
| - | [[Category: | + | </StructureSection> |
| + | [[Category: Escherichia virus Lambda]] | ||
| + | [[Category: Large Structures]] | ||
[[Category: RCSB PDB Molecule of the Month]] | [[Category: RCSB PDB Molecule of the Month]] | ||
[[Category: Transposase]] | [[Category: Transposase]] | ||
| - | [[Category: Aihara | + | [[Category: Aihara H]] |
| - | [[Category: Biswas | + | [[Category: Biswas T]] |
| - | [[Category: Ellenberger | + | [[Category: Ellenberger T]] |
| - | [[Category: Filman | + | [[Category: Filman D]] |
| - | [[Category: Landy | + | [[Category: Landy A]] |
| - | [[Category: Radman-Livaja | + | [[Category: Radman-Livaja M]] |
| - | + | ||
| - | + | ||
Current revision
Crystal structure of a lambda integrase tetramer bound to a Holliday junction
| |||||||||||