This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2psm
From Proteopedia
m (Protected "2psm" [edit=sysop:move=sysop]) |
|||
| (7 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | [[Image:2psm.png|left|200px]] | ||
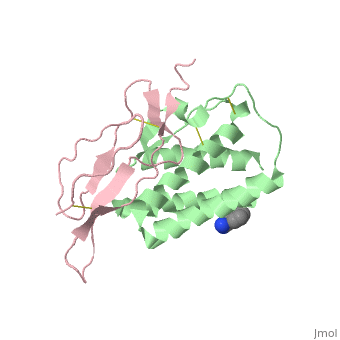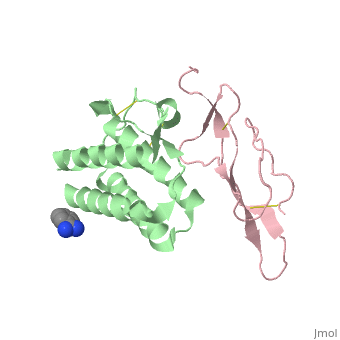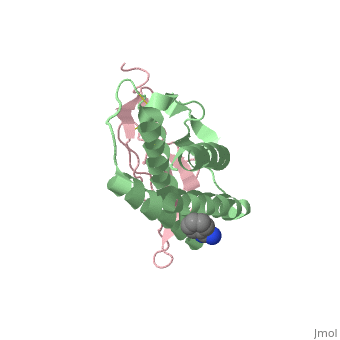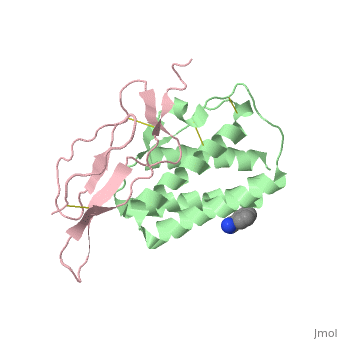
| - | + | ==Crystal structure of Interleukin 15 in complex with Interleukin 15 receptor alpha== | |
| + | <StructureSection load='2psm' size='340' side='right'caption='[[2psm]], [[Resolution|resolution]] 2.19Å' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[2psm]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Lk3_transgenic_mice Lk3 transgenic mice]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2PSM OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2PSM FirstGlance]. <br> | ||
| + | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=BEN:BENZAMIDINE'>BEN</scene></td></tr> | ||
| + | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">Il15 ([https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10090 LK3 transgenic mice]), Il15ra ([https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=10090 LK3 transgenic mice])</td></tr> | ||
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2psm FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2psm OCA], [https://pdbe.org/2psm PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2psm RCSB], [https://www.ebi.ac.uk/pdbsum/2psm PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2psm ProSAT], [https://www.topsan.org/Proteins/RSGI/2psm TOPSAN]</span></td></tr> | ||
| + | </table> | ||
| + | == Function == | ||
| + | [[https://www.uniprot.org/uniprot/IL15_MOUSE IL15_MOUSE]] Cytokine that stimulates the proliferation of T-lymphocytes. Stimulation by IL-15 requires interaction of IL-15 with components of IL-2R, including IL-2R beta and probably IL-2R gamma but not IL-2R alpha. [[https://www.uniprot.org/uniprot/I15RA_MOUSE I15RA_MOUSE]] High-affinity receptor for interleukin-15. Can signal both in cis and trans where IL15R from one subset of cells presents IL15 to neighboring IL2RG-expressing cells. Signal transduction seems to involve STAT3, STAT5, STAT6, JAK2 and SYK. Expression of different isoforms may alter or interfere with signal transduction.<ref>PMID:12734349</ref> <ref>PMID:17947230</ref> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/ps/2psm_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=2psm ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
| + | Interleukin (IL)-15 is a pleiotropic cytokine that plays a pivotal role in both innate and adaptive immunity. IL-15 is unique among cytokines due to its participation in a trans signaling mechanism in which IL-15 receptor alpha (IL-15Ralpha) from one subset of cells presents IL-15 to neighboring IL-2Rbeta/gammac-expressing cells. Here we present the crystal structure of IL-15 in complex with the sushi domain of IL-15Ralpha. The structure reveals that the alpha receptor-binding epitope of IL-15 adopts a unique conformation, which, together with amino acid substitutions, permits specific interactions with IL-15Ralpha that account for the exceptionally high affinity of the IL-15.IL-15Ralpha complex. Interestingly, analysis of the topology of IL-15 and IL-15Ralpha at the IL-15.IL-15Ralpha interface suggests that IL-15 should be capable of participating in a cis signaling mechanism similar to that of the related cytokine IL-2. Indeed, we present biochemical data demonstrating that IL-15 is capable of efficiently signaling in cis through IL-15Ralpha and IL-2Rbeta/gammac expressed on the surface of a single cell. Based on our data we propose that cis presentation of IL-15 may be important in certain biological contexts and that flexibility of IL-15Ralpha permits IL-15 and its three receptor components to be assembled identically at the ligand-receptor interface whether IL-15 is presented in cis or trans. Finally, we have gained insights into IL-15.IL-15Ralpha.IL-2Rbeta.gammac quaternary complex assembly through the use of molecular modeling. | ||
| - | + | Crystal Structure of the interleukin-15.interleukin-15 receptor alpha complex: insights into trans and cis presentation.,Olsen SK, Ota N, Kishishita S, Kukimoto-Niino M, Murayama K, Uchiyama H, Toyama M, Terada T, Shirouzu M, Kanagawa O, Yokoyama S J Biol Chem. 2007 Dec 21;282(51):37191-204. Epub 2007 Oct 18. PMID:17947230<ref>PMID:17947230</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | <div class="pdbe-citations 2psm" style="background-color:#fffaf0;"></div> | |
| - | + | ||
==See Also== | ==See Also== | ||
| - | *[[Interleukin|Interleukin]] | + | *[[Interleukin 3D structures|Interleukin 3D structures]] |
| - | *[[Interleukin receptor|Interleukin receptor]] | + | *[[Interleukin receptor 3D structures|Interleukin receptor 3D structures]] |
| - | + | == References == | |
| - | == | + | <references/> |
| - | < | + | __TOC__ |
| - | [[Category: | + | </StructureSection> |
| - | [[Category: Kanagawa, O | + | [[Category: Large Structures]] |
| - | [[Category: Kishishita, S | + | [[Category: Lk3 transgenic mice]] |
| - | [[Category: Kukimoto-Niino, M | + | [[Category: Kanagawa, O]] |
| - | [[Category: Murayama, K | + | [[Category: Kishishita, S]] |
| - | [[Category: Olsen, S K | + | [[Category: Kukimoto-Niino, M]] |
| - | [[Category: Ota, N | + | [[Category: Murayama, K]] |
| - | [[Category: | + | [[Category: Olsen, S K]] |
| - | [[Category: Shirouzu, M | + | [[Category: Ota, N]] |
| - | [[Category: Terada, T | + | [[Category: Structural genomic]] |
| - | [[Category: Yokoyama, S | + | [[Category: Shirouzu, M]] |
| + | [[Category: Terada, T]] | ||
| + | [[Category: Yokoyama, S]] | ||
| + | [[Category: Alternative splicing]] | ||
[[Category: Cytokine]] | [[Category: Cytokine]] | ||
[[Category: Endoplasmic reticulum]] | [[Category: Endoplasmic reticulum]] | ||
| Line 37: | Line 60: | ||
[[Category: Phosphorylation]] | [[Category: Phosphorylation]] | ||
[[Category: Receptor]] | [[Category: Receptor]] | ||
| - | [[Category: Riken structural genomics/proteomics initiative]] | ||
[[Category: Rsgi]] | [[Category: Rsgi]] | ||
[[Category: Secreted]] | [[Category: Secreted]] | ||
| - | [[Category: Structural genomic]] | ||
[[Category: Sushi]] | [[Category: Sushi]] | ||
[[Category: Transmembrane]] | [[Category: Transmembrane]] | ||
Current revision
Crystal structure of Interleukin 15 in complex with Interleukin 15 receptor alpha
| |||||||||||
Categories: Large Structures | Lk3 transgenic mice | Kanagawa, O | Kishishita, S | Kukimoto-Niino, M | Murayama, K | Olsen, S K | Ota, N | Structural genomic | Shirouzu, M | Terada, T | Yokoyama, S | Alternative splicing | Cytokine | Endoplasmic reticulum | Glycoprotein | Golgi apparatus | Membrane | National project on protein structural and functional analyse | Nppsfa | Nucleus | Phosphorylation | Receptor | Rsgi | Secreted | Sushi | Transmembrane