This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hemagglutinin
From Proteopedia
(Difference between revisions)
| (19 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
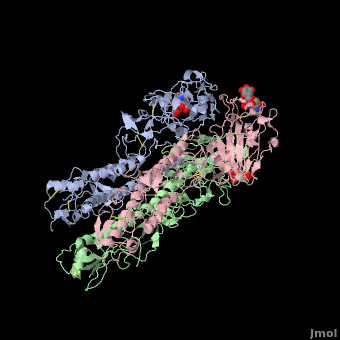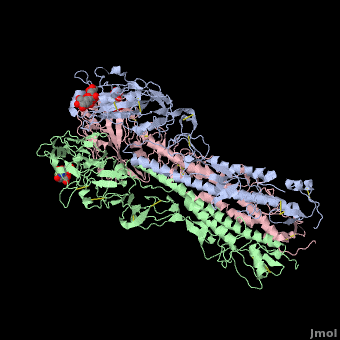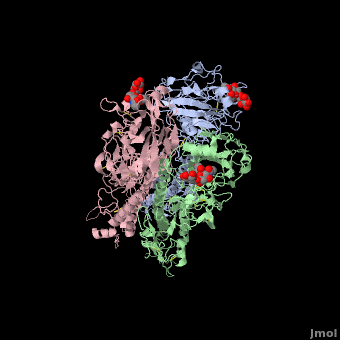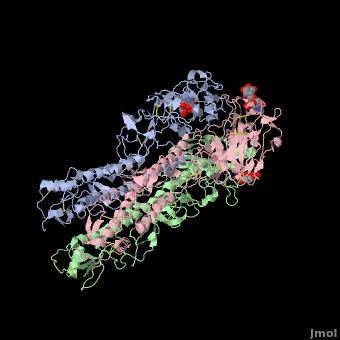
| - | <StructureSection load='' size=' | + | <StructureSection load='' size='350' side='right' scene ='Hemagglutinin/Blackground2wre/1' caption='Structure of glycosylated viral hemagglutinin trimer complex with galactose ([[2wre]])'> |
| - | [[Hemagglutinin|Hemagglutinins]] (HA) are one of the two antigenic Glycoproteins inserted into the influenza virus' membrane. There are at least 16 different forms of HA | + | == Function == |
| - | 1. On the target cell, HA | + | [[Hemagglutinin|Hemagglutinins]] (HA) are one of the two antigenic Glycoproteins inserted into the influenza virus' membrane. There are at least 16 different forms of HA antigens classified from H1 to H16. H1, H2, H3 are specific to human [[influenza]]. HAs have two main functions crucial for the [[Viral Infections|viral infection]] cycle: |
| + | 1. On the target cell, HA binds the receptor on the cell membrane which is contains sialic acid <ref>Gottschalk A. Chemistry of virus receptors. The Viruses. 1959;3:51–61.</ref> allowing the virus/cell interaction. | ||
2. HA induce the fusion between the host cell and the virus which can entry into the cytoplasm. | 2. HA induce the fusion between the host cell and the virus which can entry into the cytoplasm. | ||
| - | Because the HA are the major Antigens of the Virus, [[Antibody|Antibodies]] recognized them and for this reason HA often change. | + | Because the HA are the major Antigens of the Virus, [[Antibody|Antibodies]] recognized them and for this reason HA often change. '''Hemagglutinin-neuraminidase''' (HN) is multifunctional. It possesses both the receptor recognition and neuraminidase activities<ref>PMID:21680512</ref>. For details see [[Mumps Virus Hemagglutinin Neuraminidase Protein]]. |
For discussion of influenza hemagglutinin see<br /> | For discussion of influenza hemagglutinin see<br /> | ||
*[[Influenza hemagglutinin]]<br /> | *[[Influenza hemagglutinin]]<br /> | ||
| + | *[[Influenza]]<br /> | ||
*[[User:Michael Strong/H1N1/HA]]<br /> | *[[User:Michael Strong/H1N1/HA]]<br /> | ||
*[[User:Michael Strong/H1N1/HA/MSA]] for multiple sequence alignment. | *[[User:Michael Strong/H1N1/HA/MSA]] for multiple sequence alignment. | ||
==Hemagglutinin Structure== | ==Hemagglutinin Structure== | ||
| - | HA is an <scene name='Hemagglutinin/Homotrimer/2'>homotrimer</scene> integral membrane glycoprotein. Each monomer is synthesized like a single polypeptide chain almost 550 amino acids. This precursor is then glycosilated and cleaved into two smaller polypeptides by removal of | + | HA is an <scene name='Hemagglutinin/Homotrimer/2'>homotrimer</scene> integral membrane glycoprotein. Each monomer is synthesized like a single polypeptide chain almost 550 amino acids. This precursor is then glycosilated and cleaved into two smaller polypeptides by removal of Arginine 329; at the same time, a conformational change occurs in the monomer. |
The HA1 and HA2 subunits are covalently attached by a <scene name='SAndbox_159/Disulfide_bond/2'>disulfide bond</scene> from HA1 position 14 to HA2 position 467(*). These two chains form one monomer, and the noncovalentely association of three monomers forms one hemagglutinin molecule: (HA1+HA2)3. It is principally stabilized by packing of the alpha-helixes. All molecules are 135 Angström long. | The HA1 and HA2 subunits are covalently attached by a <scene name='SAndbox_159/Disulfide_bond/2'>disulfide bond</scene> from HA1 position 14 to HA2 position 467(*). These two chains form one monomer, and the noncovalentely association of three monomers forms one hemagglutinin molecule: (HA1+HA2)3. It is principally stabilized by packing of the alpha-helixes. All molecules are 135 Angström long. | ||
| Line 67: | Line 69: | ||
(**) The amino acid 153 lacks for the A monomer chain. | (**) The amino acid 153 lacks for the A monomer chain. | ||
| - | </StructureSection> | ||
==3D structures of hemagglutinin== | ==3D structures of hemagglutinin== | ||
| + | [[Hemagglutinin 3D structures]] | ||
| - | + | ==3D Printed Physical Models of Hemagglutinin== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | === | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | Shown below are 3D printed physical models of Hemagglutinin. The model is shown as an alpha carbon backbone, with key sidechains and domains highlighted. It has been designed with precisely embedded magnets that allow the three chains to pull apart into individual pieces. | ||
| + | |||
| + | [[Image:hemagglutinin_1_centerForBiomolecularModeling.jpg | 450px]][[Image:hemagglutinin_2_centerForBiomolecularModeling.jpg | 450px]] | ||
| + | ====The MSOE Center for BioMolecular Modeling==== | ||
| + | [[Image:CbmUniversityLogo.jpg | left | 100px]] | ||
| + | The [http://cbm.msoe.edu MSOE Center for BioMolecular Modeling] uses 3D printing technology to create physical models of protein and molecular structures, making the invisible molecular world more tangible and comprehensible. To view more protein structure models, visit our [http://cbm.msoe.edu/educationalmedia/modelgallery/ Model Gallery]. | ||
| + | </StructureSection> | ||
==Additional Resources== | ==Additional Resources== | ||
Current revision
| |||||||||||
Additional Resources
For additional information, see: Influenza
References
- ↑ Gottschalk A. Chemistry of virus receptors. The Viruses. 1959;3:51–61.
- ↑ Kim SH, Subbiah M, Samuel AS, Collins PL, Samal SK. Roles of the fusion and hemagglutinin-neuraminidase proteins in replication, tropism, and pathogenicity of avian paramyxoviruses. J Virol. 2011 Sep;85(17):8582-96. doi: 10.1128/JVI.00652-11. Epub 2011 Jun 15. PMID:21680512 doi:http://dx.doi.org/10.1128/JVI.00652-11
- ↑ Wiley DC, Skehel JJ. The structure and function of the hemagglutinin membrane glycoprotein of influenza virus. Annu Rev Biochem. 1987;56:365-94. PMID:3304138 doi:http://dx.doi.org/10.1146/annurev.bi.56.070187.002053
- ↑ Liu J, Stevens DJ, Haire LF, Walker PA, Coombs PJ, Russell RJ, Gamblin SJ, Skehel JJ. Structures of receptor complexes formed by hemagglutinins from the Asian Influenza pandemic of 1957. Proc Natl Acad Sci U S A. 2009 Oct 6;106(40):17175-80. Epub 2009 Sep 28. PMID:19805083
- ↑ Cell Binding protein in Avian Influenza; Jack Cerchiara,'06 and Brendan Holsberry, 07;http://biology.kenyon.edu/BMB/Chime2/2005/Cerchiara-Holsberry/FRAMES/start.htm
- ↑ 6.0 6.1 Knossow M, Skehel JJ. Variation and infectivity neutralization in influenza. Immunology. 2006 Sep;119(1):1-7. PMID:16925526 doi:http://dx.doi.org/10.1111/j.1365-2567.2006.02421.x
- ↑ See Influenza in Wikipedia.
Proteopedia Page Contributors and Editors (what is this?)
Julien Madouasse, Michal Harel, David Canner, Mark Hoelzer, Ann Taylor, Alexander Berchansky, Joel L. Sussman